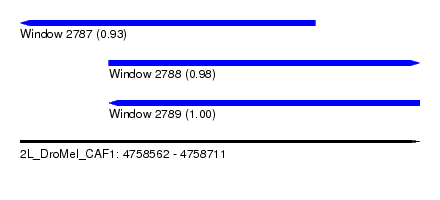

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,758,562 – 4,758,711 |
| Length | 149 |
| Max. P | 0.999944 |

| Location | 4,758,562 – 4,758,672 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 94.58 |
| Mean single sequence MFE | -26.29 |
| Consensus MFE | -22.36 |
| Energy contribution | -22.00 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.930121 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4758562 110 - 22407834 CCCAUUUGCUUUGUAACACUUACACUCGGAUGGCUUACGUAAGCCAUUUGCCUGGCCAU--AUUUUAUUUAUGCCUCAUGUGCGCUUAGUUAUUCUGGGUUUUUGUGGUAAU .((((..((((.(((((.....((..((((((((((....))))))))))..))(((((--((..............))))).))...)))))...))))....)))).... ( -26.44) >DroSec_CAF1 206871 110 - 1 CCCAUUUGCUUUGUAACACUUACACUCGGAUGGCUUACGUAAGCCAUUUGCCUGGCCAU--AUUUUAUUUAUGCCUCAUGUGCGCUUAGUUAUUCUGGGUUUUUGUGGUAAU .((((..((((.(((((.....((..((((((((((....))))))))))..))(((((--((..............))))).))...)))))...))))....)))).... ( -26.44) >DroSim_CAF1 183793 110 - 1 CCCAUUUGCUUUGUAACACUUACACUCGGAUGGCUUACGUAAGCCAUUUGCCUGGCCAU--AUUUUAUUUAUGCCUCAUGUGCGCUUAGUUAUUCUGGGUUUUUGUGGUAAU .((((..((((.(((((.....((..((((((((((....))))))))))..))(((((--((..............))))).))...)))))...))))....)))).... ( -26.44) >DroEre_CAF1 195556 112 - 1 CCUAUUUGCUUUGUAACACUUACACUCGAAUGGCUUACGUAAGCCAUUUGCUUGGCCAUAUAUUUUAUUUAUGCCUCAUGGGCGCUUAGCUAUUCUGGGUUUUCGUGGUAAU ......................((..((((((((((....))))))))))..))(((((.............(((.....)))((((((.....))))))....)))))... ( -27.10) >DroYak_CAF1 193557 110 - 1 CCUCUUUGCUUUGUAACACUUACACUCGAAUGGCUUACGUAAGCCAUUUGCUUAGCCAU--AUUUUAUUUAUGCCUCAUGUGCGUUUAGCUACGCUGGGUUUUUGUGGUAAU ...........((((.....))))..((((((((((....))))))))))....(((((--(..........((((..(((((.....)).)))..))))...))))))... ( -25.02) >consensus CCCAUUUGCUUUGUAACACUUACACUCGGAUGGCUUACGUAAGCCAUUUGCCUGGCCAU__AUUUUAUUUAUGCCUCAUGUGCGCUUAGUUAUUCUGGGUUUUUGUGGUAAU .......(((.((((.....))))..((((((((((....))))))))))...)))...............((((.((.....((((((.....))))))...)).)))).. (-22.36 = -22.00 + -0.36)



| Location | 4,758,595 – 4,758,711 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 91.23 |
| Mean single sequence MFE | -26.22 |
| Consensus MFE | -23.80 |
| Energy contribution | -23.56 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.983232 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4758595 116 + 22407834 AUGAGGCAUAAAUAAAAU--AUGGCCAGGCAAAUGGCUUACGUAAGCCAUCCGAGUGUAAGUGUUACAAAGCAAAUGGGAUAAUGGACU-GACUGCCUGGCAGAGAUAAAAGUAAUCUA ......((((.......)--)))((((((((..((((((....))))))((((..(.((..((((....))))..)).)....))))..-...))))))))..((((.......)))). ( -29.60) >DroSec_CAF1 206904 116 + 1 AUGAGGCAUAAAUAAAAU--AUGGCCAGGCAAAUGGCUUACGUAAGCCAUCCGAGUGUAAGUGUUACAAAGCAAAUGGGAUAAUGGACU-GACUGCCUGGCAGAGAUAAAAGUAAUCUA ......((((.......)--)))((((((((..((((((....))))))((((..(.((..((((....))))..)).)....))))..-...))))))))..((((.......)))). ( -29.60) >DroSim_CAF1 183826 116 + 1 AUGAGGCAUAAAUAAAAU--AUGGCCAGGCAAAUGGCUUACGUAAGCCAUCCGAGUGUAAGUGUUACAAAGCAAAUGGGAUAAUGGACU-GACUGCCUGGCAGAGAUAAAAGUAAUCUA ......((((.......)--)))((((((((..((((((....))))))((((..(.((..((((....))))..)).)....))))..-...))))))))..((((.......)))). ( -29.60) >DroEre_CAF1 195589 114 + 1 AUGAGGCAUAAAUAAAAUAUAUGGCCAAGCAAAUGGCUUACGUAAGCCAUUCGAGUGUAAGUGUUACAAAGCAAAUAGGAUAUA-----GGAUGGCCUGGCAGGGAUAAAAGUAAUCUA .....((..........(((..(((((......)))))...))).(((((((..((((...((((....))))......)))).-----)))))))...))..((((.......)))). ( -21.80) >DroYak_CAF1 193590 117 + 1 AUGAGGCAUAAAUAAAAU--AUGGCUAAGCAAAUGGCUUACGUAAGCCAUUCGAGUGUAAGUGUUACAAAGCAAAGAGGAUAUAGGUUAGGAUGGCAUGGCAGGGAUAAAAGUAAUCUA ....(.(((.........--...(((((((.....))))).))..((((((((..((((.....))))...)......(((....))).)))))))))).)..((((.......)))). ( -20.50) >consensus AUGAGGCAUAAAUAAAAU__AUGGCCAGGCAAAUGGCUUACGUAAGCCAUCCGAGUGUAAGUGUUACAAAGCAAAUGGGAUAAUGGACU_GACUGCCUGGCAGAGAUAAAAGUAAUCUA .......................(((((((....(((((....)))))((((...(((............)))....)))).............)))))))..((((.......)))). (-23.80 = -23.56 + -0.24)


| Location | 4,758,595 – 4,758,711 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 91.23 |
| Mean single sequence MFE | -23.52 |
| Consensus MFE | -23.16 |
| Energy contribution | -23.20 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.98 |
| SVM decision value | 4.73 |
| SVM RNA-class probability | 0.999944 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4758595 116 - 22407834 UAGAUUACUUUUAUCUCUGCCAGGCAGUC-AGUCCAUUAUCCCAUUUGCUUUGUAACACUUACACUCGGAUGGCUUACGUAAGCCAUUUGCCUGGCCAU--AUUUUAUUUAUGCCUCAU .((((.......))))..(((((((....-.....................((((.....))))...(((((((((....))))))))))))))))(((--(.......))))...... ( -26.80) >DroSec_CAF1 206904 116 - 1 UAGAUUACUUUUAUCUCUGCCAGGCAGUC-AGUCCAUUAUCCCAUUUGCUUUGUAACACUUACACUCGGAUGGCUUACGUAAGCCAUUUGCCUGGCCAU--AUUUUAUUUAUGCCUCAU .((((.......))))..(((((((....-.....................((((.....))))...(((((((((....))))))))))))))))(((--(.......))))...... ( -26.80) >DroSim_CAF1 183826 116 - 1 UAGAUUACUUUUAUCUCUGCCAGGCAGUC-AGUCCAUUAUCCCAUUUGCUUUGUAACACUUACACUCGGAUGGCUUACGUAAGCCAUUUGCCUGGCCAU--AUUUUAUUUAUGCCUCAU .((((.......))))..(((((((....-.....................((((.....))))...(((((((((....))))))))))))))))(((--(.......))))...... ( -26.80) >DroEre_CAF1 195589 114 - 1 UAGAUUACUUUUAUCCCUGCCAGGCCAUCC-----UAUAUCCUAUUUGCUUUGUAACACUUACACUCGAAUGGCUUACGUAAGCCAUUUGCUUGGCCAUAUAUUUUAUUUAUGCCUCAU ..................(((((((.....-----................((((.....))))...(((((((((....))))))))))))))))....................... ( -22.90) >DroYak_CAF1 193590 117 - 1 UAGAUUACUUUUAUCCCUGCCAUGCCAUCCUAACCUAUAUCCUCUUUGCUUUGUAACACUUACACUCGAAUGGCUUACGUAAGCCAUUUGCUUAGCCAU--AUUUUAUUUAUGCCUCAU ...............................................(((.((((.....))))..((((((((((....))))))))))...)))(((--(.......))))...... ( -14.30) >consensus UAGAUUACUUUUAUCUCUGCCAGGCAGUC_AGUCCAUUAUCCCAUUUGCUUUGUAACACUUACACUCGGAUGGCUUACGUAAGCCAUUUGCCUGGCCAU__AUUUUAUUUAUGCCUCAU ..................(((((((..........................((((.....))))...(((((((((....))))))))))))))))....................... (-23.16 = -23.20 + 0.04)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:09:51 2006