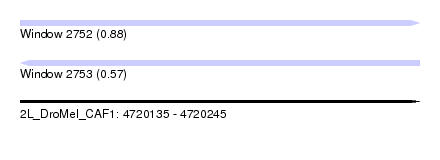

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,720,135 – 4,720,245 |
| Length | 110 |
| Max. P | 0.878407 |

| Location | 4,720,135 – 4,720,245 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 83.83 |
| Mean single sequence MFE | -31.53 |
| Consensus MFE | -22.92 |
| Energy contribution | -25.03 |
| Covariance contribution | 2.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.878407 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4720135 110 + 22407834 GCACAAAAUAUGGACAGGCUAUUUGGGCUGCAGAGGUUUUGUAGCCUGGCUUCUGGACCACUGAGUACUGUCAGCCAAUAAAGUGUUUUUCU------UGUUUUUGCUGAAAUAUC (((.((((((.(((.((((.((((.(((((((((((((....))))).((((..(.....).)))).))).)))))....)))))))).)))------)))))))))......... ( -28.70) >DroSec_CAF1 148488 102 + 1 GCACAAAAUAUGGACAGGCUAUUAGAGCUGCAGAGGUUUA--------------UGGCCACUGAGUACUGUCAGCCAAUAAAGAGUUUUUCUUUCCAUUGUUUGUGCUGAAAUAUC (((((((..(((((..((((..(((.(((.(((.(((...--------------..))).)))))).)))..))))....(((((...))))))))))..)))))))......... ( -33.30) >DroSim_CAF1 144540 102 + 1 GCACAAAAUAUGGACAGGCUAUUAGGGCUGCAGAGGUUUA--------------UGGCCACUGAGUACUGUCAGCCAAUAAAGUGUUUUUCUUUCCAUUGUUUGUGCUGAAAUAUC (((((((..(((((..((((..(((..((.(((.(((...--------------..))).)))))..)))..))))....(((.......))))))))..)))))))......... ( -32.60) >consensus GCACAAAAUAUGGACAGGCUAUUAGGGCUGCAGAGGUUUA______________UGGCCACUGAGUACUGUCAGCCAAUAAAGUGUUUUUCUUUCCAUUGUUUGUGCUGAAAUAUC (((((((...((((..((((..(((..((.(((.((((.................)))).)))))..)))..))))....(((.......)))))))...)))))))......... (-22.92 = -25.03 + 2.11)


| Location | 4,720,135 – 4,720,245 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 83.83 |
| Mean single sequence MFE | -24.97 |
| Consensus MFE | -16.03 |
| Energy contribution | -18.37 |
| Covariance contribution | 2.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.568311 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4720135 110 - 22407834 GAUAUUUCAGCAAAAACA------AGAAAAACACUUUAUUGGCUGACAGUACUCAGUGGUCCAGAAGCCAGGCUACAAAACCUCUGCAGCCCAAAUAGCCUGUCCAUAUUUUGUGC ...............(((------(((...(((..((((((((((.(((......((((.((........)))))).......))))))))..)))))..))).....)))))).. ( -21.82) >DroSec_CAF1 148488 102 - 1 GAUAUUUCAGCACAAACAAUGGAAAGAAAAACUCUUUAUUGGCUGACAGUACUCAGUGGCCA--------------UAAACCUCUGCAGCUCUAAUAGCCUGUCCAUAUUUUGUGC .........(((((((..(((((((((.....))))....(((((..((..(((((.((...--------------....)).))).))..))..)))))..)))))..))))))) ( -27.50) >DroSim_CAF1 144540 102 - 1 GAUAUUUCAGCACAAACAAUGGAAAGAAAAACACUUUAUUGGCUGACAGUACUCAGUGGCCA--------------UAAACCUCUGCAGCCCUAAUAGCCUGUCCAUAUUUUGUGC .........(((((((..((((((((.......)))....(((((..((..(((((.((...--------------....)).))).))..))..)))))..)))))..))))))) ( -25.60) >consensus GAUAUUUCAGCACAAACAAUGGAAAGAAAAACACUUUAUUGGCUGACAGUACUCAGUGGCCA______________UAAACCUCUGCAGCCCUAAUAGCCUGUCCAUAUUUUGUGC .........(((((((..(((((..(((......)))...(((((..((..(((((.((.....................)).))).))..))..)))))..)))))..))))))) (-16.03 = -18.37 + 2.33)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:09:18 2006