
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,660,141 – 4,660,236 |
| Length | 95 |
| Max. P | 0.965225 |

| Location | 4,660,141 – 4,660,236 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 91.57 |
| Mean single sequence MFE | -35.96 |
| Consensus MFE | -27.40 |
| Energy contribution | -27.64 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553462 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4660141 95 + 22407834 AAGGCGCUUCCUGCCGCUCCAAAGUGGCCCGCUGUGGCGUUUGUUUGCCACACGCAAUCAAUGGAUGUGGAUGUGGUUAGGG-----AUUGGUUUGGAAU (((.((.((((((((((((((.........(((((((((......))))))).)).(((....))).)))).)))).)))))-----).)).)))..... ( -33.20) >DroSec_CAF1 96375 94 + 1 AAGGCGCUUCCUGCCGCUCCAAAGUGGCCCGCUGUGGCGUUUGUUUGCCACACGCAAUCAAUGGUUGUGGAUGUGGUUAGGG-----AUUGGGUUGGAA- ..(((.(.((((((((((....))))))(((((((((((......)))))))((((((.....))))))...))))...)))-----)..).)))....- ( -36.20) >DroSim_CAF1 88052 95 + 1 AAGGCGCUUCCUGCCGCUCCAAAGUGGCCCGCUGUGGCGUUUGUUUGCCACACGCAAUCAAUGGUUGUGGAUGUGGUUAGGG-----AUUGGGUUGGAAU ..(((.(.((((((((((....))))))(((((((((((......)))))))((((((.....))))))...))))...)))-----)..).)))..... ( -36.20) >DroEre_CAF1 99974 100 + 1 AAGGCGCUUCCUGCCGAACCAAAGUGGCCCGCUGUGGCGUUUGUUUGCCACACGCAAUCAAUGGAUGUGGAUGUGGCUAGGGAGCGGAUUGGGUUGGGUU ....((((((((((((..(((.........(((((((((......))))))).)).(((....))).)))...)))).)))))))).............. ( -37.10) >DroYak_CAF1 100166 100 + 1 AAGGCGCUUCCUGCCGAACCAAAGUGGCCCGCUGUGGCGUUUGUUUGCCACACGCAAUCACUGGAUGUGGAUGUGGUUAAGGUUUGGAUUGGGUUGGGUU ..(((.(..(((.(((((((....(((((((((((((((......))))))).))..((((.....))))..).))))).)))))))...)))..).))) ( -37.10) >consensus AAGGCGCUUCCUGCCGCUCCAAAGUGGCCCGCUGUGGCGUUUGUUUGCCACACGCAAUCAAUGGAUGUGGAUGUGGUUAGGG_____AUUGGGUUGGAAU .((.(((.(((.(((((......)))))..(((((((((......))))))).)).............))).))).))...................... (-27.40 = -27.64 + 0.24)

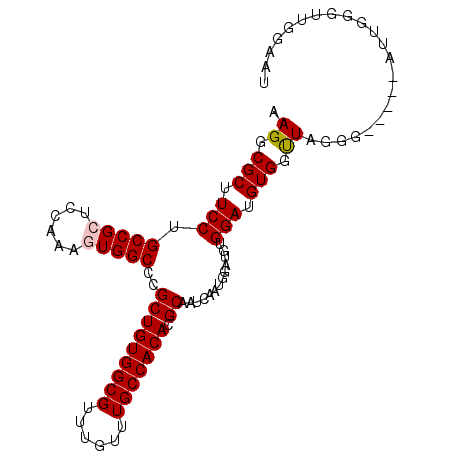

| Location | 4,660,141 – 4,660,236 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 91.57 |
| Mean single sequence MFE | -26.72 |
| Consensus MFE | -22.82 |
| Energy contribution | -22.66 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.965225 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4660141 95 - 22407834 AUUCCAAACCAAU-----CCCUAACCACAUCCACAUCCAUUGAUUGCGUGUGGCAAACAAACGCCACAGCGGGCCACUUUGGAGCGGCAGGAAGCGCCUU .(((((((.....-----(((............((.....))...((.((((((........)))))))))))....))))))).(((.(....)))).. ( -26.10) >DroSec_CAF1 96375 94 - 1 -UUCCAACCCAAU-----CCCUAACCACAUCCACAACCAUUGAUUGCGUGUGGCAAACAAACGCCACAGCGGGCCACUUUGGAGCGGCAGGAAGCGCCUU -((((((......-----(((............((.....))...((.((((((........))))))))))).....)))))).(((.(....)))).. ( -23.90) >DroSim_CAF1 88052 95 - 1 AUUCCAACCCAAU-----CCCUAACCACAUCCACAACCAUUGAUUGCGUGUGGCAAACAAACGCCACAGCGGGCCACUUUGGAGCGGCAGGAAGCGCCUU .((((((......-----(((............((.....))...((.((((((........))))))))))).....)))))).(((.(....)))).. ( -24.20) >DroEre_CAF1 99974 100 - 1 AACCCAACCCAAUCCGCUCCCUAGCCACAUCCACAUCCAUUGAUUGCGUGUGGCAAACAAACGCCACAGCGGGCCACUUUGGUUCGGCAGGAAGCGCCUU ..............((((.(((.(((...................((.((((((........))))))))(((((.....))))))))))).)))).... ( -33.70) >DroYak_CAF1 100166 100 - 1 AACCCAACCCAAUCCAAACCUUAACCACAUCCACAUCCAGUGAUUGCGUGUGGCAAACAAACGCCACAGCGGGCCACUUUGGUUCGGCAGGAAGCGCCUU ............(((................(((.....)))..(((.((((((........)))))).((((((.....))))))))))))........ ( -25.70) >consensus AUUCCAACCCAAU_____CCCUAACCACAUCCACAUCCAUUGAUUGCGUGUGGCAAACAAACGCCACAGCGGGCCACUUUGGAGCGGCAGGAAGCGCCUU ........((((......(((............((.....))...((.((((((........))))))))))).....))))...(((.(....)))).. (-22.82 = -22.66 + -0.16)

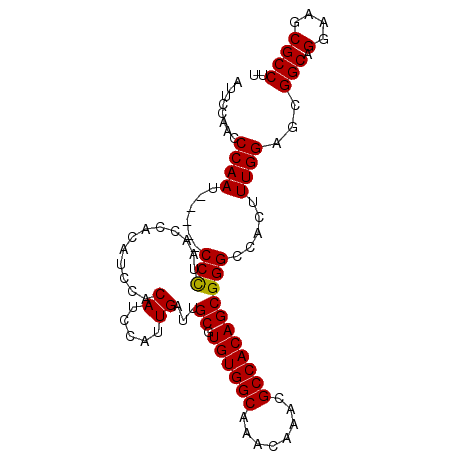
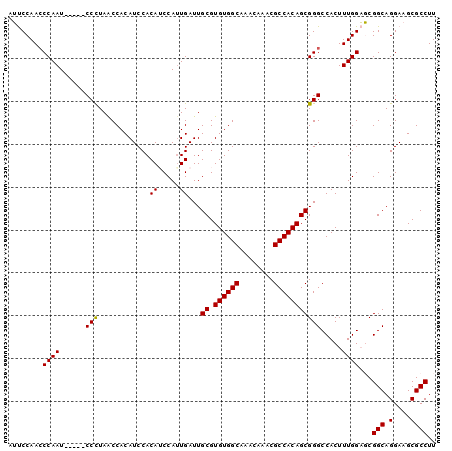
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:08:58 2006