| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,595,250 – 4,595,346 |
| Length | 96 |
| Max. P | 0.985229 |

| Location | 4,595,250 – 4,595,346 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 77.94 |
| Mean single sequence MFE | -22.39 |
| Consensus MFE | -12.44 |
| Energy contribution | -12.67 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.56 |
| SVM decision value | 2.00 |
| SVM RNA-class probability | 0.985229 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
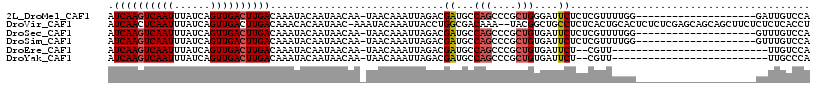
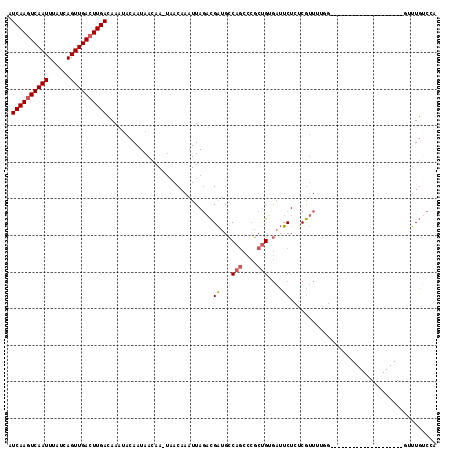
>2L_DroMel_CAF1 4595250 96 - 22407834 AUCAAGUCAAUUUAUCAGUUGACUUGACAAAUACAAUAACAA-UAACAAAUUAGACGAUGCCAGCCCGCUGGGAUUCUCUCGUUUUGG--------------------GAUUGUCCA .((((((((((......))))))))))...........((((-(..((((..(((.(((.((((....)))).))).)))...)))).--------------------.)))))... ( -30.50) >DroVir_CAF1 41502 114 - 1 AUCAACUCAAUUUAUCAGUUGACUUGACAAACACAAUAAC-AAAUACAAAUUACCUGGCGACAAA--UACGGCUGCCUCUCACUGCACUCUCUCGAGCAGCAGCUUCUCUCUCACCU .((((.(((((......))))).)))).............-...............((.((....--...((((((..(((.............)))..))))))......)).)). ( -19.84) >DroSec_CAF1 40122 96 - 1 AUCAAGUCAAUUUAUCAGUUGACUUGACAAAUACAAUAACAA-UAACAAAUUAGACGAUGCCAGCCCGCUGUGAUUCUCUCGUUUUGG--------------------GUUUGUCCA .((((((((((......))))))))))...........((((-...((((..(((.(((..(((....)))..))).)))...)))).--------------------..))))... ( -22.50) >DroSim_CAF1 37733 96 - 1 AUCAAGUCAAUUUAUCAGUUGACUUGACAAAUACAAUAACAA-UAACAAAUUAGACGAUGCCAGCCCGCUGUGAUUCUCUCGUUUUGG--------------------GUUUGUCCA .((((((((((......))))))))))...........((((-...((((..(((.(((..(((....)))..))).)))...)))).--------------------..))))... ( -22.50) >DroEre_CAF1 42816 88 - 1 AUCAAGUCAAUUUAUCAGUUGACUUGACAAAUACAAUAACAA-UAACAAAUUAGACGAUGCCAGCCCGCUGUGAUUCU--CGUU--------------------------UUGUCCA .((((((((((......))))))))))...............-..(((((...((.(((..(((....)))..))).)--)..)--------------------------))))... ( -20.10) >DroYak_CAF1 43275 88 - 1 AUCAAGUCAAUUUAUCAGUUGACUUGACAAAUACAAUAACAA-UAACAAAUUAGACGAUGCCAGCCCGCUGUGAUUCU--CGUU--------------------------UUGCCCA .((((((((((......))))))))))...............-...((((...((.(((..(((....)))..))).)--)..)--------------------------))).... ( -18.90) >consensus AUCAAGUCAAUUUAUCAGUUGACUUGACAAAUACAAUAACAA_UAACAAAUUAGACGAUGCCAGCCCGCUGUGAUUCUCUCGUUUUGG____________________GUUUGUCCA .((((((((((......)))))))))).............................((...(((....)))....))........................................ (-12.44 = -12.67 + 0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:08:25 2006