
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 518,161 – 518,267 |
| Length | 106 |
| Max. P | 0.942799 |

| Location | 518,161 – 518,267 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 80.31 |
| Mean single sequence MFE | -31.80 |
| Consensus MFE | -16.45 |
| Energy contribution | -17.12 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.26 |
| Mean z-score | -3.12 |
| Structure conservation index | 0.52 |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.942799 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
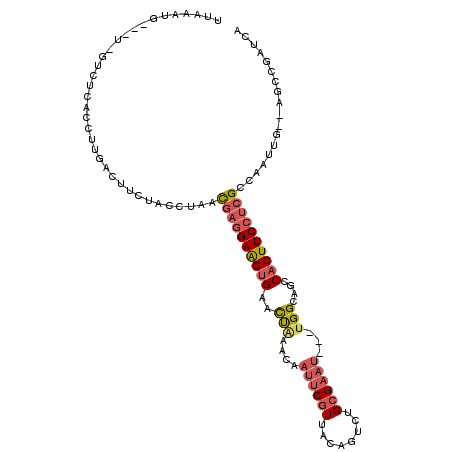
>2L_DroMel_CAF1 518161 106 + 22407834 UGAAAUGGAAU-GUCUCGCCUUGACUUCUACCUAACGAGGAAGUGAACUACCCAAUUCGUUACAGUCUGCGAAU---UGGCAGUCACUUCCUCGCAAAUUG--AUCCGAUCA .((..(((((.-(((.......)))))))).....(((((((((((.((..(((((((((........))))))---))).))))))))))))).......--......)). ( -37.30) >DroSec_CAF1 63727 103 + 1 UUAAGUG---U-GUCUCACCUUGACUUCUACCUAACGAGGAAGUGAACUAAACAAUUCGUUACAGUCUGCGAAU---UGGCAGCCACUUCCUCGCCAAUUG--AGCCGAUCA (((((.(---(-(...))))))))...........((((((((((..((...((((((((........))))))---))..)).)))))))))).......--......... ( -29.60) >DroSim_CAF1 65852 103 + 1 UUAAAUG---U-GUCUCACCUUGACUUCUACCUAACGAGGAAGUGAACUAAACAAUUCGUUACAGUCUGCGAAU---UGGCAGCCACUUCCUCGCCAAUUG--AGCCGAUCA ((((.((---.-(((.......)))..........((((((((((..((...((((((((........))))))---))..)).)))))))))).)).)))--)........ ( -28.30) >DroEre_CAF1 67234 103 + 1 UCAAAUG---U-GCCUCACCUUGACUUCUACCCAACGAGGAAGUGUAUUGCCCAAUUCGUUACAGUCUGCGAAC---UGGCAGCCACUUCCUCGCCAAUUG--AGCCGAUCA ((((.((---.-((.((.....))............(((((((((..(((((...(((((........))))).---.))))).))))))))))))).)))--)........ ( -30.20) >DroYak_CAF1 66256 103 + 1 UUAAAUG---U-GUCUCACCUUGAGUUCUGCCUAACGAGGAAGUGUACCAAACAAUUCGUCACAGUCUGCGAAU---UGGAAGCCACUUCCUCGCCAAUCA--AGCUGAUCA .......---.-...(((.((((((((......)))(((((((((.......((((((((........))))))---)).....))))))))).....)))--)).)))... ( -30.30) >DroAna_CAF1 81404 109 + 1 UUAGAGG---CGGUUAUUCCUUUUUUUUUUAAUAAUUAAGAGGGGUAUCUUCAAAUUCGUCACAGUCUGGGUCUCUGUGGCAGCCACUUCCCUGCCACUUGAGAGCCGAUCA .....((---(((.((((((((((..((.....))..)))))))))).........)))))...(((.((.((((.(((((((........)))))))..)))).))))).. ( -35.10) >consensus UUAAAUG___U_GUCUCACCUUGACUUCUACCUAACGAGGAAGUGAACUAAACAAUUCGUUACAGUCUGCGAAU___UGGCAGCCACUUCCUCGCCAAUUG__AGCCGAUCA ...................................((((((((((..(((....((((((........))))))...)))....)))))))))).................. (-16.45 = -17.12 + 0.67)


| Location | 518,161 – 518,267 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 80.31 |
| Mean single sequence MFE | -35.07 |
| Consensus MFE | -18.59 |
| Energy contribution | -19.32 |
| Covariance contribution | 0.73 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.81 |
| Structure conservation index | 0.53 |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.909632 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
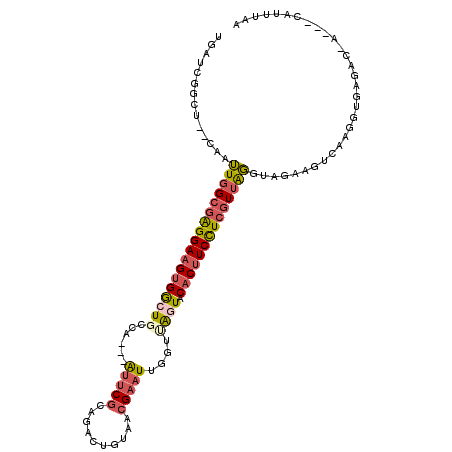
>2L_DroMel_CAF1 518161 106 - 22407834 UGAUCGGAU--CAAUUUGCGAGGAAGUGACUGCCA---AUUCGCAGACUGUAACGAAUUGGGUAGUUCACUUCCUCGUUAGGUAGAAGUCAAGGCGAGAC-AUUCCAUUUCA .....((((--(.((((((((((((((((((((((---(((((..........)))))).))))).))))))))))).))))).)).(((.......)))-..)))...... ( -39.80) >DroSec_CAF1 63727 103 - 1 UGAUCGGCU--CAAUUGGCGAGGAAGUGGCUGCCA---AUUCGCAGACUGUAACGAAUUGUUUAGUUCACUUCCUCGUUAGGUAGAAGUCAAGGUGAGAC-A---CACUUAA .....((((--(..((((((((((((((((((.((---(((((..........)))))))..))).)))))))))))))))...).)))).(((((....-.---))))).. ( -37.00) >DroSim_CAF1 65852 103 - 1 UGAUCGGCU--CAAUUGGCGAGGAAGUGGCUGCCA---AUUCGCAGACUGUAACGAAUUGUUUAGUUCACUUCCUCGUUAGGUAGAAGUCAAGGUGAGAC-A---CAUUUAA .....((((--(..((((((((((((((((((.((---(((((..........)))))))..))).)))))))))))))))...).)))).(((((....-.---))))).. ( -34.60) >DroEre_CAF1 67234 103 - 1 UGAUCGGCU--CAAUUGGCGAGGAAGUGGCUGCCA---GUUCGCAGACUGUAACGAAUUGGGCAAUACACUUCCUCGUUGGGUAGAAGUCAAGGUGAGGC-A---CAUUUGA .....(.((--((.(..(((((((((((..(((((---(((((..........)))))).))))...)))))))))))..).............)))).)-.---....... ( -38.44) >DroYak_CAF1 66256 103 - 1 UGAUCAGCU--UGAUUGGCGAGGAAGUGGCUUCCA---AUUCGCAGACUGUGACGAAUUGUUUGGUACACUUCCUCGUUAGGCAGAACUCAAGGUGAGAC-A---CAUUUAA ...(((.((--((((((((((((((((((((..((---(((((((.....)).)))))))...))).)))))))))))))).......))))).)))...-.---....... ( -38.61) >DroAna_CAF1 81404 109 - 1 UGAUCGGCUCUCAAGUGGCAGGGAAGUGGCUGCCACAGAGACCCAGACUGUGACGAAUUUGAAGAUACCCCUCUUAAUUAUUAAAAAAAAAAGGAAUAACCG---CCUCUAA .....(((((((..(((((((........))))))).)))).((...........(((...((((......))))....)))..........)).......)---))..... ( -22.00) >consensus UGAUCGGCU__CAAUUGGCGAGGAAGUGGCUGCCA___AUUCGCAGACUGUAACGAAUUGGUUAGUACACUUCCUCGUUAGGUAGAAGUCAAGGUGAGAC_A___CAUUUAA ..............((((((((((((((((((......(((((..........)))))....)))).))))))))))))))............................... (-18.59 = -19.32 + 0.73)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:27:52 2006