
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,338,797 – 4,338,887 |
| Length | 90 |
| Max. P | 0.549198 |

| Location | 4,338,797 – 4,338,887 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 83.05 |
| Mean single sequence MFE | -27.32 |
| Consensus MFE | -16.44 |
| Energy contribution | -16.25 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.549198 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
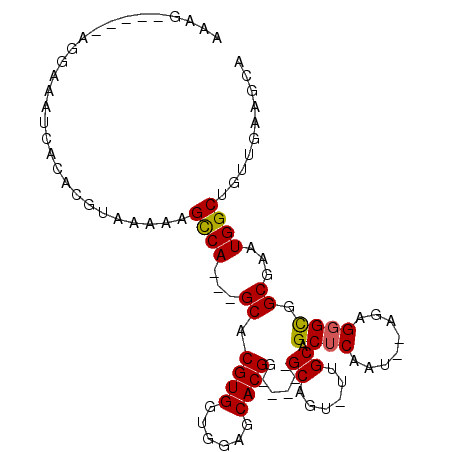
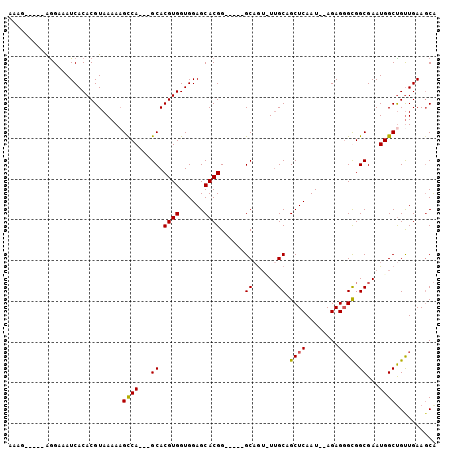
>2L_DroMel_CAF1 4338797 90 - 22407834 AUAG-----AGGAAAUCACACGUAAAAAGCCA---GCACGUGGUGGAGCACGG-----GCAGUUUUGCAGCUCAAU--AGAGUGUGGCGAAUGGCCGUUGAAGCA ....-----...................((((---((.((((......))))(-----((...(((((.((((...--.))))...)))))..)))))))..)). ( -25.50) >DroSec_CAF1 7714 89 - 1 AAAG-----AGGAAAUCACACGUAAAAUGCCA---GCACGUGGUGGAGCACGG-----GCCGU-UUGCAGCUCAAU--AGAGGGCGGCGAAUGGCUGUUGAAGCA ....-----..................(((((---((.((((......))))(-----(((((-((((.((((...--...)))).))))))))))))))..))) ( -31.20) >DroSim_CAF1 8329 92 - 1 AAAG-----AGGAAAUCACACGUAAAAAGCCAGCAGCACGUGGUGGAGCACGG-----GCAGU-UUGCAGCUCAAU--AGAGGGCGGCGAAUGGCUGUUGAAUCA ....-----.....................(((((((.((((......)))).-----...((-((((.((((...--...)))).)))))).)))))))..... ( -27.40) >DroEre_CAF1 7774 102 - 1 AAAAAGGAAGAGAAAUCACACGUAAAAAGUCA---GCACGUGGUGGAGCACGGCAACUGCAAACUGGCAGCUCAAUAAAGAGGGUGGCGAAUGGCUGCUGAAGCG .........((....))............(((---(((.((.((...((.((.(..((((......))))(((......)))).))))..)).)))))))).... ( -25.20) >consensus AAAG_____AGGAAAUCACACGUAAAAAGCCA___GCACGUGGUGGAGCACGG_____GCAGU_UUGCAGCUCAAU__AGAGGGCGGCGAAUGGCUGUUGAAGCA ............................((((...((.((((......))))......((......)).((((........)))).))...)))).......... (-16.44 = -16.25 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:06:40 2006