
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,333,676 – 4,333,776 |
| Length | 100 |
| Max. P | 0.792627 |

| Location | 4,333,676 – 4,333,776 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 74.70 |
| Mean single sequence MFE | -29.38 |
| Consensus MFE | -19.52 |
| Energy contribution | -21.08 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.792627 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
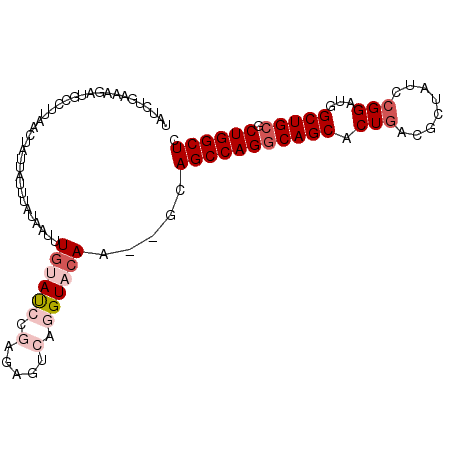
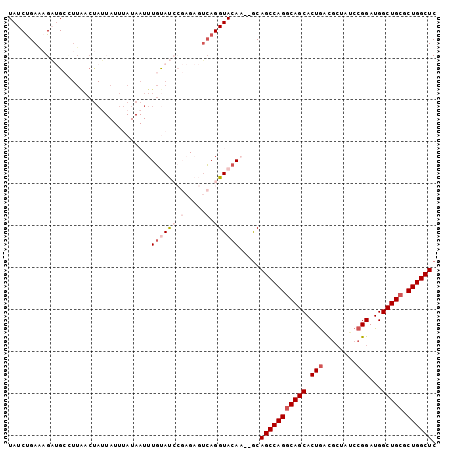
>2L_DroMel_CAF1 4333676 100 + 22407834 UAUCUGAAAGAUGCGUUAACUAUUAUUUAUAAUUUGUAUCCGAGAGUCAGGUACAA--GCAGCCAGGCAGCACUAUUGCUAUCCGGAUGGCUGCGCUGGCUC (((((((..((((((...................))))))(....))))))))...--..(((((((((((.(.(((........))))))))).)))))). ( -27.81) >DroSec_CAF1 2647 100 + 1 UAUCUGAAAGAUGCCUUAACUAUUAUUUAUAAUUUGUAUCCGAGAGUCAGGUACAA--GCAGCCAGGCAGCACUGACACUAUCCGGAUGGCUGCGCUGGCUC (((((((..(((((...((.(((.....))).)).)))))(....))))))))...--..(((((((((((.(((........)))...))))).)))))). ( -29.50) >DroSim_CAF1 2570 100 + 1 UAUCUGAAAGAUGCCUUGACUAUUAUUUAUAAUUUGUAUCCGAGAGUCAGGUACAA--ACAGCCAGGCAGCACUGACGCUAUCCGGAUGGCUGCGCUGGCUC ...........(((((.((((......(((.....)))......)))))))))...--..(((((((((((.(((........)))...))))).)))))). ( -30.70) >DroEre_CAF1 2631 84 + 1 UAUCCGAGUGUAGGGUUAACCAU----------UUGCAC------AGCAGGUACAC--GCAGCCAGGCAGCACUGGCGGUAUCCGGAUGGCUGCGCUGGCUC .....(.((((.((.....))((----------((((..------.))))))))))--.)(((((((((((.((((......))))...))))).)))))). ( -32.90) >DroYak_CAF1 2799 79 + 1 UAUCUGAAUGUCGAU-------G----------UUCCAC------AUGAGGUACACAAGCAGCCAGCCAGCACUGACGCUAUCCGGAUGGCUGCGCUGGCUC (((((..((((.(..-------.----------..).))------)).))))).......(((((((((((.(((........)))...)))).))))))). ( -26.00) >consensus UAUCUGAAAGAUGCCUUAACUAUUAUUUAUAAUUUGUAUCCGAGAGUCAGGUACAA__GCAGCCAGGCAGCACUGACGCUAUCCGGAUGGCUGCGCUGGCUC ..................................((((((.(.....).)))))).....(((((((((((.(((........)))...))))).)))))). (-19.52 = -21.08 + 1.56)

| Location | 4,333,676 – 4,333,776 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 74.70 |
| Mean single sequence MFE | -28.14 |
| Consensus MFE | -17.88 |
| Energy contribution | -19.28 |
| Covariance contribution | 1.40 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.566813 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
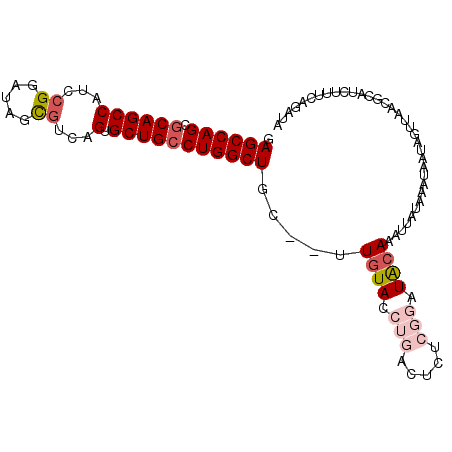
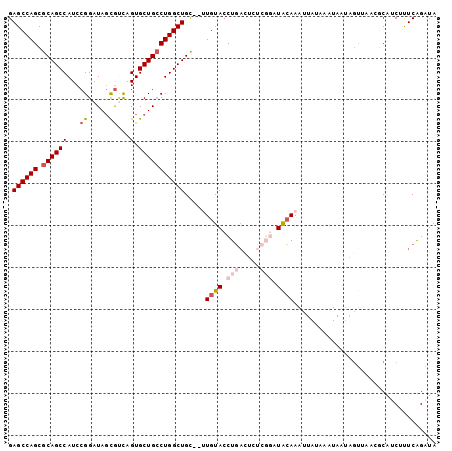
>2L_DroMel_CAF1 4333676 100 - 22407834 GAGCCAGCGCAGCCAUCCGGAUAGCAAUAGUGCUGCCUGGCUGC--UUGUACCUGACUCUCGGAUACAAAUUAUAAAUAAUAGUUAACGCAUCUUUCAGAUA ......(((((((((...((.(((((....))))))))))))))--(((((.(((.....))).)))))...................))(((.....))). ( -28.10) >DroSec_CAF1 2647 100 - 1 GAGCCAGCGCAGCCAUCCGGAUAGUGUCAGUGCUGCCUGGCUGC--UUGUACCUGACUCUCGGAUACAAAUUAUAAAUAAUAGUUAAGGCAUCUUUCAGAUA ..(((...(((((((...((.((((......)))))))))))))--(((((.(((.....))).)))))..................)))(((.....))). ( -28.90) >DroSim_CAF1 2570 100 - 1 GAGCCAGCGCAGCCAUCCGGAUAGCGUCAGUGCUGCCUGGCUGU--UUGUACCUGACUCUCGGAUACAAAUUAUAAAUAAUAGUCAAGGCAUCUUUCAGAUA ..(((...(((((((...((.(((((....))))))))))))))--(((((.(((.....))).)))))..................)))(((.....))). ( -28.30) >DroEre_CAF1 2631 84 - 1 GAGCCAGCGCAGCCAUCCGGAUACCGCCAGUGCUGCCUGGCUGC--GUGUACCUGCU------GUGCAA----------AUGGUUAACCCUACACUCGGAUA .((((((((((((((..(((.(((.....))))))..)))))))--))((((.....------))))..----------.)))))...((.......))... ( -28.90) >DroYak_CAF1 2799 79 - 1 GAGCCAGCGCAGCCAUCCGGAUAGCGUCAGUGCUGGCUGGCUGCUUGUGUACCUCAU------GUGGAA----------C-------AUCGACAUUCAGAUA ..(((((.(((((((.((((...((....)).)))).)))))))))).))..((.((------((.(..----------.-------..).))))..))... ( -26.50) >consensus GAGCCAGCGCAGCCAUCCGGAUAGCGUCAGUGCUGCCUGGCUGC__UUGUACCUGACUCUCGGAUACAAAUUAUAAAUAAUAGUUAACGCAUCUUUCAGAUA .((((((.((((((...((.....))...).))))))))))).....((((.(((.....))).)))).................................. (-17.88 = -19.28 + 1.40)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:06:37 2006