| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,282,777 – 4,282,877 |
| Length | 100 |
| Max. P | 0.998037 |

| Location | 4,282,777 – 4,282,877 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 69.38 |
| Mean single sequence MFE | -24.96 |
| Consensus MFE | -7.58 |
| Energy contribution | -7.58 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.30 |
| SVM decision value | 2.99 |
| SVM RNA-class probability | 0.998037 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
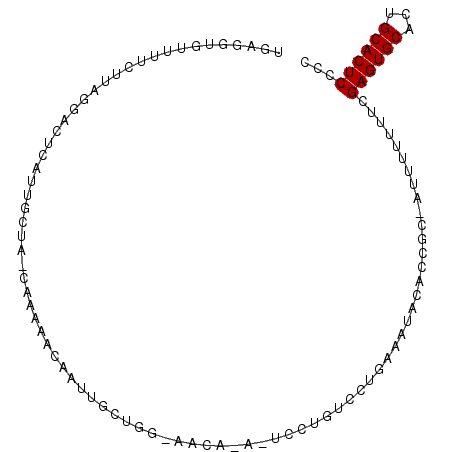
>2L_DroMel_CAF1 4282777 100 + 22407834 UUAGGUGUUUUCUUAGGUCUCAUUUCUA-CAAGAACAAUUGCUGGAAACAUAUUCCUGGCCUGAAUUACACCGC-AUUUGUUUCGAGUGCACUGCACUCCCC ...(((((....(((((((...((((((-(((......))).)))))).........)))))))...)))))..-.........((((((...))))))... ( -26.30) >DroSec_CAF1 37154 101 + 1 UUAGGUGUUUUCUUAGGACUCAUUGCUACCAUUAACAAUUGCUGGCAACACAUUCCUGCCCAGAAAUACACCGC-AUUUUUUUCGAGUGCACUGCACUCCCC ...(((((((((((((((....((((((.((........)).)))))).....)))))...))))).)))))..-.........((((((...))))))... ( -29.00) >DroSim_CAF1 38555 101 + 1 UGAGGUGUUUUCUUAGGACUCAUUGCUAGCAAUAACAAUUGCUGGCAACACAAUCCUGUCCAGAAAUACAACGC-AUUUGUUUCGAGUGCACUGCACUCCCC .((((((((((..(((((....(((((((((((....))))))))))).....)))))....)))))))...((-(((((...)))))))......)))... ( -33.20) >DroEre_CAF1 33163 84 + 1 UGAAGGGUUUUCUCUAGUUUU-CUCUUA-CAAACACAAUUGCC---------------UCAUGAAAGGCGCCCU-GUUUUUUUCGAGUGCACUGCACUCCCC ...(((((........((((.-......-.))))......(((---------------(......)))))))))-.........((((((...))))))... ( -20.30) >DroYak_CAF1 37848 86 + 1 UAAGCGUGUUUCUUUCGCUUUAGUGUUA-CACACACAAUCGCC---------------ACAUGAACUGCACCAAGGUUUUUUUAGAGUGCACUGCACUCCCC ...(((.((((.....((..(.((((..-...)))).)..)).---------------....)))))))...............((((((...))))))... ( -16.00) >consensus UGAGGUGUUUUCUUAGGACUCAUUGCUA_CAAAAACAAUUGCUGG_AACA_A_UCCUGUCCUGAAAUACACCGC_AUUUUUUUCGAGUGCACUGCACUCCCC ....................................................................................((((((...))))))... ( -7.58 = -7.58 + 0.00)


| Location | 4,282,777 – 4,282,877 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 69.38 |
| Mean single sequence MFE | -27.24 |
| Consensus MFE | -8.19 |
| Energy contribution | -10.39 |
| Covariance contribution | 2.20 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.30 |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.937572 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
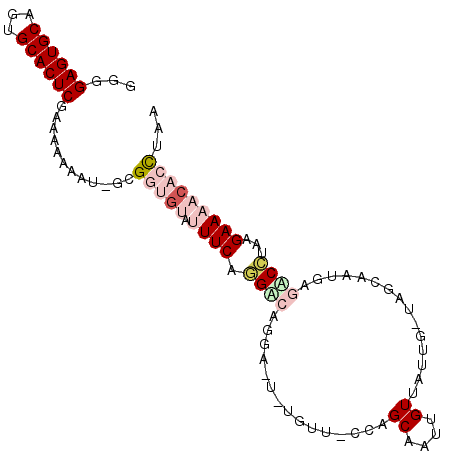
>2L_DroMel_CAF1 4282777 100 - 22407834 GGGGAGUGCAGUGCACUCGAAACAAAU-GCGGUGUAAUUCAGGCCAGGAAUAUGUUUCCAGCAAUUGUUCUUG-UAGAAAUGAGACCUAAGAAAACACCUAA (((((((((...))))))...((((.(-(((((.(.....).))).((((.....)))).))).))))(((..-(....)..)))))).............. ( -25.80) >DroSec_CAF1 37154 101 - 1 GGGGAGUGCAGUGCACUCGAAAAAAAU-GCGGUGUAUUUCUGGGCAGGAAUGUGUUGCCAGCAAUUGUUAAUGGUAGCAAUGAGUCCUAAGAAAACACCUAA ...((((((...)))))).........-..(((((.(((((....((((...((((((((...........)))))))).....)))).))))))))))... ( -33.20) >DroSim_CAF1 38555 101 - 1 GGGGAGUGCAGUGCACUCGAAACAAAU-GCGUUGUAUUUCUGGACAGGAUUGUGUUGCCAGCAAUUGUUAUUGCUAGCAAUGAGUCCUAAGAAAACACCUCA (((((((((...)))))).........-....(((.(((((....((((((.((((((.((((((....)))))).)))))))))))).)))))))).))). ( -33.30) >DroEre_CAF1 33163 84 - 1 GGGGAGUGCAGUGCACUCGAAAAAAAC-AGGGCGCCUUUCAUGA---------------GGCAAUUGUGUUUG-UAAGAG-AAAACUAGAGAAAACCCUUCA (((((((((...)))))).....((((-(.((.(((((....))---------------)))..)).))))).-......-..............))).... ( -22.90) >DroYak_CAF1 37848 86 - 1 GGGGAGUGCAGUGCACUCUAAAAAAACCUUGGUGCAGUUCAUGU---------------GGCGAUUGUGUGUG-UAACACUAAAGCGAAAGAAACACGCUUA ..(((((((...)))))))...........((((...(((...(---------------.((....((((...-..))))....)).)..)))...)))).. ( -21.00) >consensus GGGGAGUGCAGUGCACUCGAAAAAAAU_GCGGUGUAUUUCAGGACAGGA_U_UGUU_CCAGCAAUUGUUAUUG_UAGCAAUGAGACCUAAGAAAACACCUAA ...((((((...))))))............(((((.((((.((((...............((....))...............))))...)))))))))... ( -8.19 = -10.39 + 2.20)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:06:16 2006