
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,042,021 – 4,042,113 |
| Length | 92 |
| Max. P | 0.729701 |

| Location | 4,042,021 – 4,042,113 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 84.93 |
| Mean single sequence MFE | -15.52 |
| Consensus MFE | -9.60 |
| Energy contribution | -9.64 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.729701 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
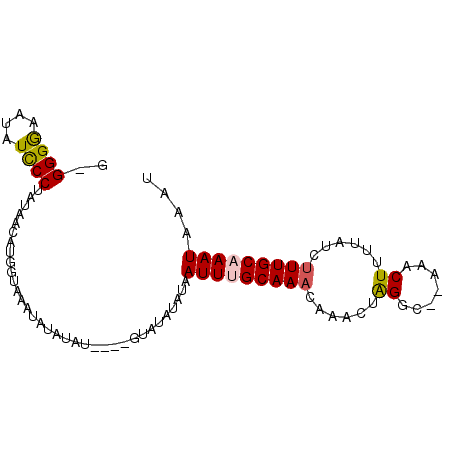
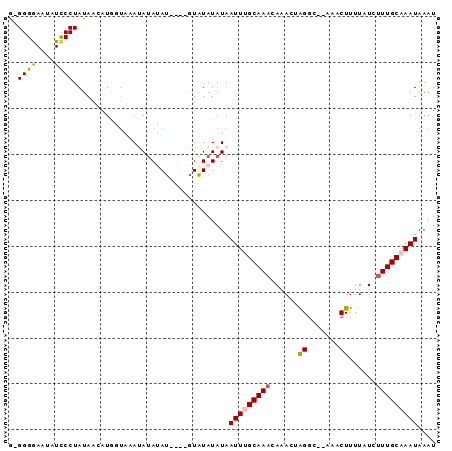
>2L_DroMel_CAF1 4042021 92 - 22407834 G-GGGGAAAAUCCCUAUAACAUGGUAAAUAUAUACAUAUAUGUAUAUAAUUUGCAAACAAACUAGGCACAAACUUUUAUCUUUGCAAAUAAAU .-((((.....)))).............((((((((....))))))))(((((((((......((.......))......))))))))).... ( -19.50) >DroSec_CAF1 174979 87 - 1 GCGGGGAAUAUCCCUAUAACAUGGUAAAUAUAUAU----GUAUAUAUAAUUGGCAAACAAACUAGGC--AAACUUUUAUCUUUGCAAAUAAAU ..((((.....))))......((.(((.((((((.----...)))))).))).))..........((--(((........)))))........ ( -14.00) >DroSim_CAF1 179528 87 - 1 GCGGGGAAUAUCCCUAUAGCAUGGUAAAUAUAUAU----GUAUAUAUAAUUGGCAAACAAACUAGGC--AAACUUUUAUCUUUGCAAAUAAAU ((((((.....))))...((.((((...((((((.----...)))))).(((.....))))))).))--..............))........ ( -15.10) >DroEre_CAF1 182896 84 - 1 A-GGGAAAUAUCCCUAUAACAUG-UGCACAUA-AU----GUAUAUAUAAUUUGCAAACAAACAAGGC--AAACUUUUAUCUUUGCAAAUAAAU (-((((....))))).....(((-((((....-.)----))))))...(((((((((.....(((..--...))).....))))))))).... ( -18.80) >DroYak_CAF1 187140 80 - 1 A-GGGAAAUAUUCCUAUAACAUA----ACA-A-AU----GUAUAUAUAAUUUGCAAACAAACAGGCC--AAACUUUUAUCAUUGCAAAUAAAU (-((((....)))))...((((.----...-.-))----)).......((((((((......((...--...)).......)))))))).... ( -10.22) >consensus G_GGGGAAUAUCCCUAUAACAUGGUAAAUAUAUAU____GUAUAUAUAAUUUGCAAACAAACUAGGC__AAACUUUUAUCUUUGCAAAUAAAU ..((((....))))..................................(((((((((......((.......))......))))))))).... ( -9.60 = -9.64 + 0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:05 2006