
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,734,794 – 3,734,887 |
| Length | 93 |
| Max. P | 0.844215 |

| Location | 3,734,794 – 3,734,887 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 89.87 |
| Mean single sequence MFE | -14.70 |
| Consensus MFE | -11.44 |
| Energy contribution | -11.92 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.844215 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
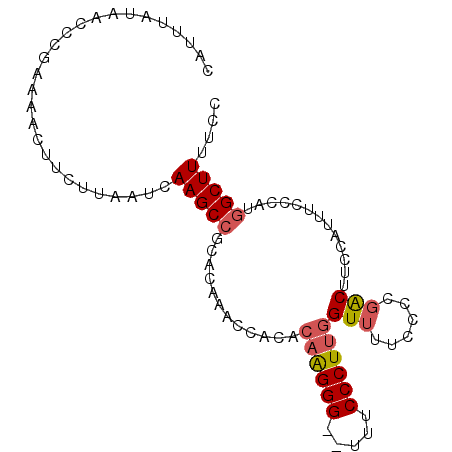
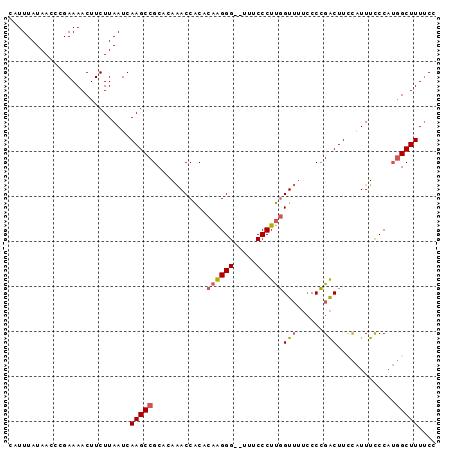
>2L_DroMel_CAF1 3734794 93 + 22407834 CAUUUAUAACCCGAAAACUUCUUAAUCAAGCCGCACAAACCACACAAGGG--UUUCCCUUGGUUUUCCCCGACUUCCAUUUCCCAUGGCUUUUCC ...........................((((((...........((((((--...))))))(((......)))............)))))).... ( -13.90) >DroSec_CAF1 197015 93 + 1 CAUUUAUAACCCGAAAACUUCUUAAUCAAGCCGCACAAACCACACAAGGG--UUUCCCUUGGUUUUCCCCGACUUCCAUUUCCCAUGGCUUUUCC ...........................((((((...........((((((--...))))))(((......)))............)))))).... ( -13.90) >DroSim_CAF1 204188 93 + 1 CAUUUAUAACCCGAAAACUUCUUAAUCAAGCCGCACAAACCACACAAGGG--UUUCCCUUGGUUUUCCCCGACUUCCAUUUCCCAUGGCUUUUCC ...........................((((((...........((((((--...))))))(((......)))............)))))).... ( -13.90) >DroYak_CAF1 210609 93 + 1 CAUUUAUAACCCGAAAACUUCUUAAUCAAGCCGCACAAACCACACAAGGG--UUUCCCUUGGUUUUCCCCGACUUUCAUUUCCCAUGGCUUUCCC ...........................((((((...........((((((--...))))))(((......)))............)))))).... ( -13.90) >DroAna_CAF1 222130 90 + 1 CAUUUAUAACCCGAAAACUUCUUAAUCAAGCCGCACAGACCGCAUUGGGGGUUUUCCCUCCGU----CCGGGCUUAUGUUUUCCC-AGCUUUUCC ............((((((.........(((((((.......))...(((((.....)))))..----...)))))..))))))..-......... ( -17.90) >consensus CAUUUAUAACCCGAAAACUUCUUAAUCAAGCCGCACAAACCACACAAGGG__UUUCCCUUGGUUUUCCCCGACUUCCAUUUCCCAUGGCUUUUCC ...........................(((((............((((((.....))))))(((......))).............))))).... (-11.44 = -11.92 + 0.48)

| Location | 3,734,794 – 3,734,887 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 89.87 |
| Mean single sequence MFE | -23.20 |
| Consensus MFE | -19.28 |
| Energy contribution | -19.32 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.826364 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
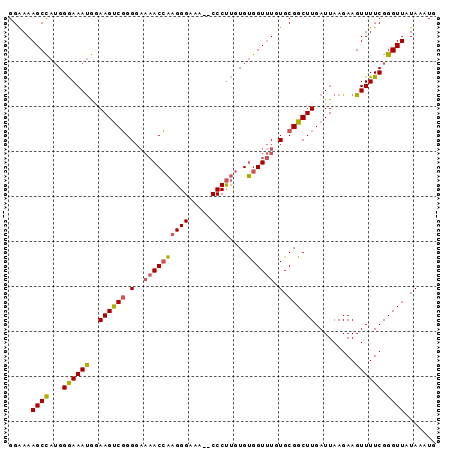
>2L_DroMel_CAF1 3734794 93 - 22407834 GGAAAAGCCAUGGGAAAUGGAAGUCGGGGAAAACCAAGGGAAA--CCCUUGUGUGGUUUGUGCGGCUUGAUUAAGAAGUUUUCGGGUUAUAAAUG .(((((.((((.....))))((((((.(.(((((((((((...--)))))).))..))).).))))))..........)))))............ ( -23.40) >DroSec_CAF1 197015 93 - 1 GGAAAAGCCAUGGGAAAUGGAAGUCGGGGAAAACCAAGGGAAA--CCCUUGUGUGGUUUGUGCGGCUUGAUUAAGAAGUUUUCGGGUUAUAAAUG .(((((.((((.....))))((((((.(.(((((((((((...--)))))).))..))).).))))))..........)))))............ ( -23.40) >DroSim_CAF1 204188 93 - 1 GGAAAAGCCAUGGGAAAUGGAAGUCGGGGAAAACCAAGGGAAA--CCCUUGUGUGGUUUGUGCGGCUUGAUUAAGAAGUUUUCGGGUUAUAAAUG .(((((.((((.....))))((((((.(.(((((((((((...--)))))).))..))).).))))))..........)))))............ ( -23.40) >DroYak_CAF1 210609 93 - 1 GGGAAAGCCAUGGGAAAUGAAAGUCGGGGAAAACCAAGGGAAA--CCCUUGUGUGGUUUGUGCGGCUUGAUUAAGAAGUUUUCGGGUUAUAAAUG .....((((...((((((..((((((.(.(((((((((((...--)))))).))..))).).)))))).........)))))).))))....... ( -23.00) >DroAna_CAF1 222130 90 - 1 GGAAAAGCU-GGGAAAACAUAAGCCCGG----ACGGAGGGAAAACCCCCAAUGCGGUCUGUGCGGCUUGAUUAAGAAGUUUUCGGGUUAUAAAUG .....((((-.(((((...(((((((((----((.(.(((......)))....).)))))...)))))).........))))).))))....... ( -22.80) >consensus GGAAAAGCCAUGGGAAAUGGAAGUCGGGGAAAACCAAGGGAAA__CCCUUGUGUGGUUUGUGCGGCUUGAUUAAGAAGUUUUCGGGUUAUAAAUG .....((((...((((((..((((((.(..((((((((((.....))))....)))))).).)))))).........)))))).))))....... (-19.28 = -19.32 + 0.04)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:01:46 2006