
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,688,756 – 3,688,856 |
| Length | 100 |
| Max. P | 0.628671 |

| Location | 3,688,756 – 3,688,856 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.68 |
| Mean single sequence MFE | -18.60 |
| Consensus MFE | -12.74 |
| Energy contribution | -12.60 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.03 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.628671 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
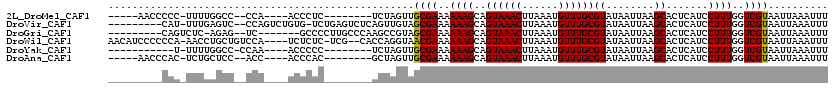
>2L_DroMel_CAF1 3688756 100 - 22407834 -----AACCCCC-UUUUGGCC--CCA----ACCCUC--------UCUAGUUGCGAAAAAAGCAGUAAACUUAAAUGUUUGCGUAUAAUUAAGCACUCAUCCUUUGGUCGUAAUUAAAUUU -----.......-..(((...--.))----).....--------..(((((((((..((((..((((((......))))))((........)).......))))..)))))))))..... ( -18.30) >DroVir_CAF1 215106 107 - 1 ---------CAU-UUUGAGUC--CCAGUCUGUG-UCUGAGUCUCAGUUGUAGCGAAAAAAGCAGUAAACUUAAAUGUUUGCGUAUAAUUAAGCACUCAUCCUUUGGUCGUAAUUAAAUUU ---------...-........--((((.....(-..(((((((.(((((((((.......)).((((((......)))))).))))))).)).)))))..).)))).............. ( -20.90) >DroGri_CAF1 190256 101 - 1 ---------CAGUCUC-AGAG--UC-------GCCCCUUGCCCAAGCCGUAGCGAAAAAAGCAGUAAACUUAAAUGUUUGCGUAUAAUUAAGCACUCAUCCUUUGGUCGUAAUUAAAUUU ---------..(.(.(-((((--((-------((..(..((....)).)..))))........((((((......))))))...................)))))).)............ ( -15.80) >DroWil_CAF1 238504 112 - 1 AACAUCCCCCCA-AACCUGCUGUCCA----UCUCUC-UCG--CACCAGGUAACGAAAAAAGCAGUAAACUUAAAUGUUUGCGUAUAAUUAAGCACUCAUCCUUUGGUCGUAAUUAAAUUU ............-.(((((.(((...----......-..)--)).))))).((((..((((..((((((......))))))((........)).......))))..)))).......... ( -19.60) >DroYak_CAF1 161537 95 - 1 -----------U-UUUUGGCC-CCAA----ACCCCC--------UCUAGUUGCGAAAAAAGCAGUAAACUUAAAUGUUUGCGUAUAAUUAAGCACUCAUCCUUUGGUCGUAAUUAAAUUU -----------.-.((((...-.)))----).....--------..(((((((((..((((..((((((......))))))((........)).......))))..)))))))))..... ( -19.20) >DroAna_CAF1 167694 100 - 1 -----AACCCAC-UCUGCUCC--ACC----ACCCAC--------GCUAGUUGCGAAAAAAGCAGUAAACUUAAAUGUUUGCGUAUAAUUAAGCACUCAUCCUUUGGUCGUAAUUAAAUUU -----.......-........--...----......--------..(((((((((..((((..((((((......))))))((........)).......))))..)))))))))..... ( -17.80) >consensus _________CAC_UUUGAGCC__CCA____ACCCUC________ACUAGUAGCGAAAAAAGCAGUAAACUUAAAUGUUUGCGUAUAAUUAAGCACUCAUCCUUUGGUCGUAAUUAAAUUU ...................................................((((..((((..((((((......))))))((........)).......))))..)))).......... (-12.74 = -12.60 + -0.14)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:01:21 2006