
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,666,340 – 3,666,574 |
| Length | 234 |
| Max. P | 0.971721 |

| Location | 3,666,340 – 3,666,434 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 87.30 |
| Mean single sequence MFE | -18.16 |
| Consensus MFE | -16.00 |
| Energy contribution | -16.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.68 |
| SVM RNA-class probability | 0.971721 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3666340 94 + 22407834 CUCGCUGAUUGACUUGUGUUUCCAUUUUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUCCGGAUGCUCGCGCGGGUAUUUUU ((((((((((((.(((.(((.(((.........)))..))).))).))))))).((((((.(((.......))).)).)))))))))....... ( -23.10) >DroSec_CAF1 128774 91 + 1 CUCGCUGAUUGACUUGUGUUUCCAUUUUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUUAUUCCGGCUCCUCGUGCGU---UUUUU ..((((((((((.(((.(((.(((.........)))..))).))).))))))).((((.(((((.......)))))..))))))).---..... ( -18.80) >DroEre_CAF1 131314 91 + 1 CUCGCUGAUUGACUUGUGUUUCCAUUUUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUCCUGCUCCUCGUGCU---AUUUUU .(((((((((((.(((.(((.(((.........)))..))).))).)))))).)))))...............((.......)).---...... ( -16.70) >DroYak_CAF1 138372 84 + 1 CUCGCUGAUUGACUUGUGUUUCCAUUUUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUCCUGCG-------UU---UUUUUU .(((((((((((.(((.(((.(((.........)))..))).))).)))))).)))))(((..((....)))))..-------..---...... ( -16.20) >DroAna_CAF1 143298 85 + 1 CUCGCUGAUUGACUUGUGUUUCCAUUUUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUUCUCCACCCC---------UUUCC .(((((((((((.(((.(((.(((.........)))..))).))).)))))).)))))......................---------..... ( -16.00) >consensus CUCGCUGAUUGACUUGUGUUUCCAUUUUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUCCUGCUCCUCG_GCG____UUUUU .(((((((((((.(((.(((.(((.........)))..))).))).)))))).))))).................................... (-16.00 = -16.00 + 0.00)


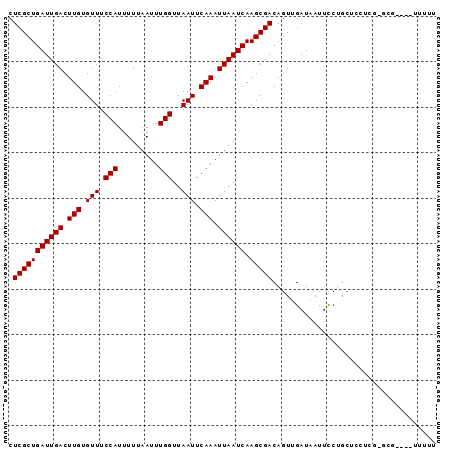
| Location | 3,666,340 – 3,666,434 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 87.30 |
| Mean single sequence MFE | -16.52 |
| Consensus MFE | -14.40 |
| Energy contribution | -14.40 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3666340 94 - 22407834 AAAAAUACCCGCGCGAGCAUCCGGAAUUAUCAACUGUCGCUUGAUUAAUUUGAAUUAACCAAAUUAAAAAUGGAAACACAAGUCAAUCAGCGAG ..........((....))..................(((((.((((.(((((((((.....)))).....((....))))))).))))))))). ( -18.90) >DroSec_CAF1 128774 91 - 1 AAAAA---ACGCACGAGGAGCCGGAAUAAUCAACUGUCGCUUGAUUAAUUUGAAUUAACCAAAUUAAAAAUGGAAACACAAGUCAAUCAGCGAG .....---.(((.((......))...............(((((.((((((((.......))))))))...((....)))))))......))).. ( -16.00) >DroEre_CAF1 131314 91 - 1 AAAAAU---AGCACGAGGAGCAGGAAUUAUCAACUGUCGCUUGAUUAAUUUGAAUUAACCAAAUUAAAAAUGGAAACACAAGUCAAUCAGCGAG ......---....((..((((((..........)))).(((((.((((((((.......))))))))...((....)))))))...))..)).. ( -16.60) >DroYak_CAF1 138372 84 - 1 AAAAAA---AA-------CGCAGGAAUUAUCAACUGUCGCUUGAUUAAUUUGAAUUAACCAAAUUAAAAAUGGAAACACAAGUCAAUCAGCGAG ......---..-------..(((..........)))(((((.((((.(((((((((.....)))).....((....))))))).))))))))). ( -15.30) >DroAna_CAF1 143298 85 - 1 GGAAA---------GGGGUGGAGAAAUUAUCAACUGUCGCUUGAUUAAUUUGAAUUAACCAAAUUAAAAAUGGAAACACAAGUCAAUCAGCGAG .....---------..(((.((.......)).))).(((((.((((.(((((((((.....)))).....((....))))))).))))))))). ( -15.80) >consensus AAAAA____AGC_CGAGGAGCAGGAAUUAUCAACUGUCGCUUGAUUAAUUUGAAUUAACCAAAUUAAAAAUGGAAACACAAGUCAAUCAGCGAG ....................................(((((.((((.(((((((((.....)))).....((....))))))).))))))))). (-14.40 = -14.40 + 0.00)



| Location | 3,666,366 – 3,666,467 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 81.75 |
| Mean single sequence MFE | -25.78 |
| Consensus MFE | -14.16 |
| Energy contribution | -15.28 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.834622 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3666366 101 + 22407834 UUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUCCGGAUGCUCGCGCGGGUAUUUUU-----UUUUCGCUGGCGCAGCGAAGUGCAUUU--GUUGCA .(((((((((.....)))))))))....(((((((.((........((((((((....))))))))..-----.((((((((...))))))))..)).))--))))). ( -30.20) >DroSec_CAF1 128800 98 + 1 UUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUUAUUCCGGCUCCUCGUGCGU---UUUUU-----UUUUCGCUGGCGCAGCGAAGUGCAUUU--GUUGCA .(((((((((.....)))))))))....((((((((((.......)))).....(((((.---.....-----.((((((((...))))))))))))).)--))))). ( -25.81) >DroEre_CAF1 131340 98 + 1 UUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUCCUGCUCCUCGUGCU---AUUUUU-----UUUUUGCUGGCGCUGCGAAGUGCAUUU--GUUGCA .(((((((((.....)))))))))....(((((((.((....((((.((.((....((.---......-----.....)).)).)).).)))...)).))--))))). ( -23.40) >DroYak_CAF1 138398 96 + 1 UUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUCCUGCG-------UU---UUUUUUGCUCGUUUUCGCUGGCGCAGCGAAGUGCAUUU--GUUGCA .(((((((((.....)))))))))....(((((((.((....((((((((-------((---......((........)).))))))).)))...)).))--))))). ( -28.50) >DroAna_CAF1 143324 94 + 1 UUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUUCUCCACCCC---------UUUCC-----UGAAGCCUGGAGCUGCAAAAUGCAUUUGUGUUGCA .(((((((((.....)))))))))....((((((...........(((((...(---------(((..-----.))))..))))).(((.....)))....)))))). ( -21.00) >consensus UUUAAUUUGGUUAAUUCAAAUUAAUCAAGCGACAGUUGAUAAUUCCUGCUCCUCG_GCG____UUUUU_____UUUUCGCUGGCGCAGCGAAGUGCAUUU__GUUGCA .(((((((((.....)))))))))....(((((....((....)).............................((((((((...)))))))).........))))). (-14.16 = -15.28 + 1.12)

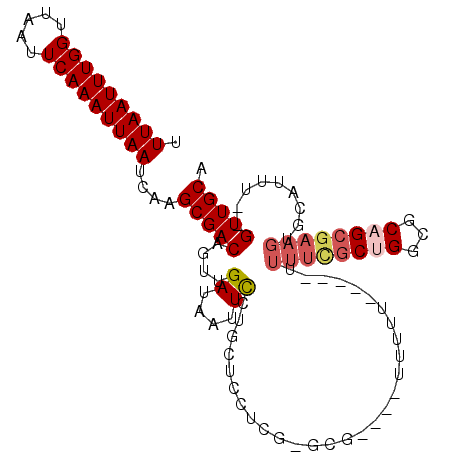

| Location | 3,666,467 – 3,666,574 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.90 |
| Mean single sequence MFE | -28.18 |
| Consensus MFE | -19.86 |
| Energy contribution | -21.10 |
| Covariance contribution | 1.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.736675 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3666467 107 - 22407834 AAACAACAAAGUGGGAAAUGCAAUGCGUCUGGACAACGGAAAGCCCAACAA-------CAUUGGAU------GCACAUGAAUAUGGAUGAGUCUGGACGCCAAUGCCAAUGCACUUUCCA ...........((((((.((((..(((((..(((........((((((...-------..))))..------)).(((........))).)))..))))).........)))).)))))) ( -28.10) >DroSec_CAF1 128898 107 - 1 AAACAACAAAGUGGGAAAUGCAAUGCGUCUGGACAACGGAAAGCCCAACAA-------CAUUGGAU------GCACAUGAAUAUGGAUGAGUCUGGACGCCAAUGCCAAUGCACUUUCGA .......((((((.(....(((..(((((..(((........((((((...-------..))))..------)).(((........))).)))..)))))...)))...).))))))... ( -27.90) >DroEre_CAF1 131438 107 - 1 AAACAACAAAGUGGGAAAUGCACUGCGUCUGGACAACGGAAAGCCCAACAA-------CAUUGGAU------GCACAUGAAUAUGGAUGAGUCUGGACGCCAAUGCCAAUGCACUCUCCA .........((((.(....(((..(((((..(((........((((((...-------..))))..------)).(((........))).)))..)))))...)))...).))))..... ( -26.10) >DroYak_CAF1 138494 114 - 1 AAACAACAAAGUGGGAAAUGCAAUGCGUCUGGACAACGGAAAGCCCAACAACAUUCAACAUUGGAU------GCACAUGAAUAUGGAUGAGUCUGGACGCCAAUGCCAAUGCACUUUCCA ...........((((((.((((..(((((..(((...((.....)).....((((((....((...------...))......)))))).)))..))))).........)))).)))))) ( -30.70) >DroAna_CAF1 143418 108 - 1 AAACAACAAAGUGGGAAGU-----GCGUCUGGACAUCAGCAACGACAACGA-------CAAUGCAGAAAUAAAUAUAUGAAUAUGUAUGAGUCUGGACUCCAAUGCCAAUGCACUUUCCA ...........((((((((-----(((((..(((.((((((.((....)).-------...)))........(((((....))))).))))))..)))....((....)))))))))))) ( -28.10) >consensus AAACAACAAAGUGGGAAAUGCAAUGCGUCUGGACAACGGAAAGCCCAACAA_______CAUUGGAU______GCACAUGAAUAUGGAUGAGUCUGGACGCCAAUGCCAAUGCACUUUCCA .......((((((.(....(((..(((((..(((..........((((............))))...........(((........))).)))..)))))...)))...).))))))... (-19.86 = -21.10 + 1.24)


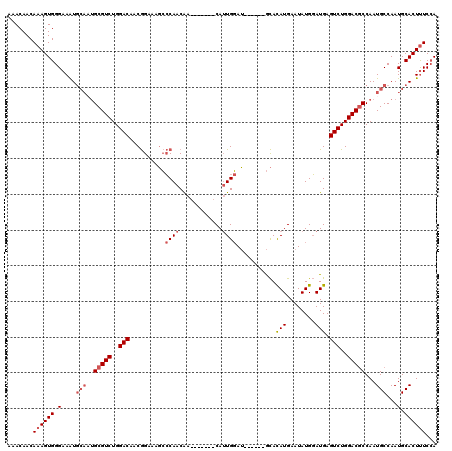
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:01:02 2006