
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,588,362 – 3,588,463 |
| Length | 101 |
| Max. P | 0.962266 |

| Location | 3,588,362 – 3,588,463 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 90.93 |
| Mean single sequence MFE | -30.61 |
| Consensus MFE | -24.85 |
| Energy contribution | -26.93 |
| Covariance contribution | 2.08 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.802113 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3588362 101 + 22407834 GUUGUUGUCUUCGCUCGAUGUUGCUUGUCUGCAGCGCCGCCAGCAACAUGGCAACAUAGCAACAGCAACAUGGACAACAGCACCAGCAGCAGAUGCACUCG ((((((((((.......((((((((.((.(((.((.......))...(((....))).)))))))))))))))))))))))....(((.....)))..... ( -35.01) >DroSec_CAF1 56076 98 + 1 GUUGUUGUCUUCGCUCGAUGUUGCUUGUCUGCAGCGCCGCCAGCAACACGGCAACAUAGCAACAGCAACAUGGACAACAGCAA---CAGCAGAUGCACUCA ((((((((((.......((((((((.((.(((.((.......)).....(....)...)))))))))))))))))))))))..---..((....))..... ( -31.91) >DroSim_CAF1 59491 98 + 1 GUUGUUGUCUUCGCUCGAUGUUGCUUGUCUGCAGCGCCGCCAGCAACACGGCAACAUAGCAACAGCAACAUGGACAACAGCAA---CAGCAGAUGCACUCG ((((((((((.......((((((((.((.(((.((.......)).....(....)...)))))))))))))))))))))))..---..((....))..... ( -31.91) >DroEre_CAF1 58169 87 + 1 -----------CGCUCGAUGUUGCUUGUCUGCAGCGCCGCCAGCAACACGGCAACAUAGCAACAGCAACAUGGACAACAGCAC---CAGCGGAUGCACUGG -----------.((((.((((((((.((.(((.((.......)).....(....)...))))))))))))).)).....)).(---(((.(....).)))) ( -25.70) >DroYak_CAF1 59013 98 + 1 GUUGUUGUCUUCGCUCGAUGUUGCUUGUCUGCAGCGCCGCCAGCAACACGGCAACAUAGCAACAGCAACAUGGCCAACAACAC---CAGCAGAUGCACUGA (((((((.....(((..((((((((.((.(((.((.......)).....(....)...)))))))))))))))))))))))..---..((....))..... ( -28.50) >consensus GUUGUUGUCUUCGCUCGAUGUUGCUUGUCUGCAGCGCCGCCAGCAACACGGCAACAUAGCAACAGCAACAUGGACAACAGCAC___CAGCAGAUGCACUCG ((((((((((.......((((((((.((.(((.((.......)).....(....)...))))))))))))))))))))))).......((....))..... (-24.85 = -26.93 + 2.08)

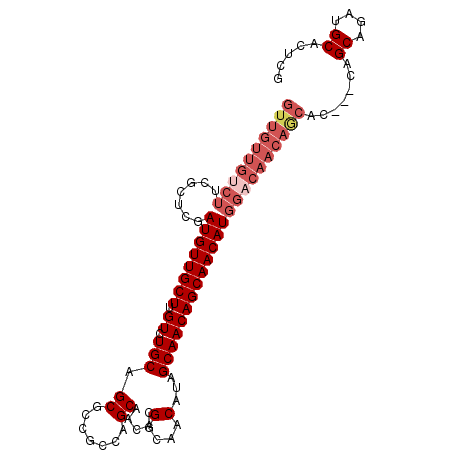

| Location | 3,588,362 – 3,588,463 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 90.93 |
| Mean single sequence MFE | -37.46 |
| Consensus MFE | -30.12 |
| Energy contribution | -30.00 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.962266 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3588362 101 - 22407834 CGAGUGCAUCUGCUGCUGGUGCUGUUGUCCAUGUUGCUGUUGCUAUGUUGCCAUGUUGCUGGCGGCGCUGCAGACAAGCAACAUCGAGCGAAGACAACAAC .....(((((.......)))))(((((((..(((((((.((((..((((((((......))))))))..))))...)))))))((....)).))))))).. ( -39.80) >DroSec_CAF1 56076 98 - 1 UGAGUGCAUCUGCUG---UUGCUGUUGUCCAUGUUGCUGUUGCUAUGUUGCCGUGUUGCUGGCGGCGCUGCAGACAAGCAACAUCGAGCGAAGACAACAAC .....((....))((---(((((.(((..(((((((((.((((..((((((((......))))))))..))))...)))))))).)..))))).))))).. ( -36.30) >DroSim_CAF1 59491 98 - 1 CGAGUGCAUCUGCUG---UUGCUGUUGUCCAUGUUGCUGUUGCUAUGUUGCCGUGUUGCUGGCGGCGCUGCAGACAAGCAACAUCGAGCGAAGACAACAAC .....((....))((---(((((.(((..(((((((((.((((..((((((((......))))))))..))))...)))))))).)..))))).))))).. ( -36.30) >DroEre_CAF1 58169 87 - 1 CCAGUGCAUCCGCUG---GUGCUGUUGUCCAUGUUGCUGUUGCUAUGUUGCCGUGUUGCUGGCGGCGCUGCAGACAAGCAACAUCGAGCG----------- ((((((....)))))---).......(..(((((((((.((((..((((((((......))))))))..))))...)))))))).)..).----------- ( -35.10) >DroYak_CAF1 59013 98 - 1 UCAGUGCAUCUGCUG---GUGUUGUUGGCCAUGUUGCUGUUGCUAUGUUGCCGUGUUGCUGGCGGCGCUGCAGACAAGCAACAUCGAGCGAAGACAACAAC (((((......))))---)(((((((.(((((((((((.((((..((((((((......))))))))..))))...)))))))).).))...))))))).. ( -39.80) >consensus CGAGUGCAUCUGCUG___GUGCUGUUGUCCAUGUUGCUGUUGCUAUGUUGCCGUGUUGCUGGCGGCGCUGCAGACAAGCAACAUCGAGCGAAGACAACAAC ....(((....((.......)).....((.((((((((.((((..((((((((......))))))))..))))...)))))))).)))))........... (-30.12 = -30.00 + -0.12)

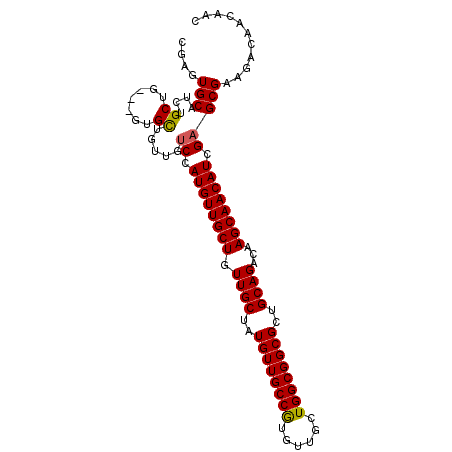

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:00:26 2006