
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,566,959 – 3,567,062 |
| Length | 103 |
| Max. P | 0.764899 |

| Location | 3,566,959 – 3,567,062 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 79.31 |
| Mean single sequence MFE | -34.74 |
| Consensus MFE | -24.63 |
| Energy contribution | -25.17 |
| Covariance contribution | 0.54 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.764899 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
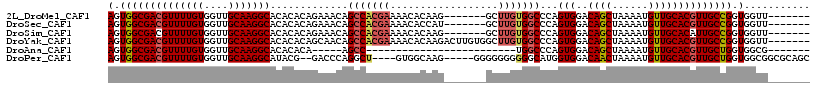
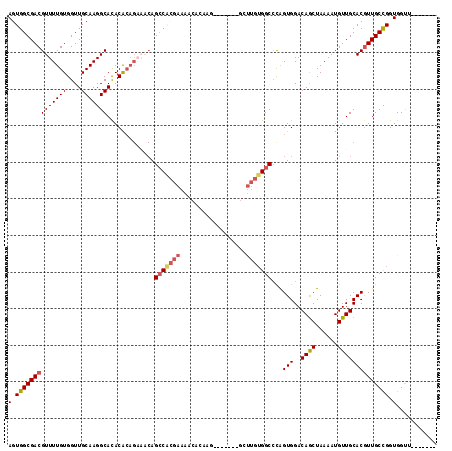
>2L_DroMel_CAF1 3566959 103 + 22407834 AGUGGCGACGUUUUGUGGUUGCAAGGCACACACAGAAACAGCCACGAAAACACAAG-------GCUUGUGGCCCAGUGGACAGCUAAAAUGUUGCACGUUGCCGGUGGUU------- (.(((((((((((((((((((.................))))))))))..(((..(-------((.....)))..)))(.((((......)))))))))))))).)....------- ( -34.93) >DroSec_CAF1 35549 103 + 1 AGUGGCGACGUUUUGUGGUUGCAAGGCACACACAGAAACAGCCACGAAAACACCAU-------GCUUGUGGCCCAGUGGACAGCUAAAAUGUUGCACGUUGCCGGUGGUU------- .(((((....(((((((..(((...)))..)))))))...))))).....((((((-------....))(((...(((..((((......)))))))...)))))))...------- ( -34.50) >DroSim_CAF1 35837 103 + 1 AGUGGCGACGUUUUGUGGUUGCAAGGCACACACAGAAACAGCCACGAAAACACAAG-------GCUUGUGGCCCAGUGGACAGCUAAAAUGUUGCACAUUGCCGGUGGUU------- .(((((....(((((((..(((...)))..)))))))...))))).....(((..(-------((.....)))(((((..((((......))))..)))))...)))...------- ( -36.50) >DroYak_CAF1 37115 110 + 1 AGUGGCGACGUUUUGUGGUUGCAAGGCACACACAGCAACAGCCACGAAAACACAAGACUUGUGGCUUGUGGCCCAGUGGACAGCUAAAAUGUUGCACGUUGCCGGUGGUU------- (.(((((((((..((((..(((...)))..))))(((((((((((((...((((.....))))..)))))))..(((.....)))....))))))))))))))).)....------- ( -42.20) >DroAna_CAF1 39396 80 + 1 AGUGGCGACGUUUUGUGGUUGCAAGGCACACACA-----AGCC-------------------------UGGCCCAGUGGACAGCUAAAAUGUUGCACGUUGCUGGUGGCG------- .(..((.((.....)).))..).((((.......-----.)))-------------------------)(.(((((..(.((((......))))....)..)))).).).------- ( -25.80) >DroPer_CAF1 38197 106 + 1 AGUGGCGACGUUUUGUGGUUGCAAGGCAUACG--GACCCAGGCU----GUGGCAAG-----GGGGGGGGGGCAUGGUGGACAACUAAAAUGUUGCACGUUGCUGGUGGCGGCGCAGC ....((((((((((((..(..(...((.....--..(((..((.----...))..)-----)).......))...)..)...)).)))))))))).((((((.....)))))).... ( -34.54) >consensus AGUGGCGACGUUUUGUGGUUGCAAGGCACACACAGAAACAGCCACGAAAACACAAG_______GCUUGUGGCCCAGUGGACAGCUAAAAUGUUGCACGUUGCCGGUGGUU_______ (.((((((((((((((....))))))).............(((((((..................)))))))...(((..((((......)))))))))))))).)........... (-24.63 = -25.17 + 0.54)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:00:09 2006