| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,414,274 – 3,414,372 |
| Length | 98 |
| Max. P | 0.937350 |

| Location | 3,414,274 – 3,414,372 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 74.50 |
| Mean single sequence MFE | -25.19 |
| Consensus MFE | -11.86 |
| Energy contribution | -12.79 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.47 |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.937350 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3414274 98 - 22407834 CUUUCUUUUUU---------UGCUUCGAUG---------GUCGCCAGCUUCUGGCAAUUUGUCCUCCCCGGCGACCCUCUGCAAA--CAGUGCAUCCAUCAACUGAUUGUUCAACUGU ..........(---------(((...((.(---------((((((.......(((.....)))......))))))).)).))))(--((((....(.(((....))).)....))))) ( -25.92) >DroAna_CAF1 6003 102 - 1 CCUUC----UAU--------UUCCACAAUAAUACACAAUUCCGACUGCCUCCGACAAUUUGUCCCCCUCGG--GCCCUCGACAAA--CAGUGCAUCUAUCAACUGAUUGUUGAACUGU .....----...--------.....(((((((.........(((..((((..(((.....)))......))--))..))).....--((((..........)))))))))))...... ( -16.90) >DroEre_CAF1 3434 107 - 1 CUCUUUUGUUUUUUUCUGUCUGCUCCGAUG---------GUCGCCAGCUUCUGGCAAUUUGUCCUCCCCGGCGACCCUCUGCAAA--CAGUGCAUCCAUCAACUGAUUGUUCCACUGC ....................(((...((.(---------((((((.......(((.....)))......))))))).)).)))..--(((((...(.(((....))).)...))))). ( -25.82) >DroYak_CAF1 3864 101 - 1 CCUUCCUGUUUU--------UGCUUCGAUG---------GUCGCCGGCUUCUGGCAAUUUGUCCUCCUCGGCGACCCUCUGCAAACACAGUGCAUCCAUCAACUGAUUGUUUCACUGU ..........((--------(((...((.(---------((((((((.....(((.....)))....))))))))).)).))))).((((((...(.(((....))).)...)))))) ( -32.10) >consensus CCUUCUUGUUUU________UGCUUCGAUG_________GUCGCCAGCUUCUGGCAAUUUGUCCUCCCCGGCGACCCUCUGCAAA__CAGUGCAUCCAUCAACUGAUUGUUCAACUGU .......................................((((((.......(((.....)))......))))))............(((((...(.(((....))).)...))))). (-11.86 = -12.79 + 0.94)

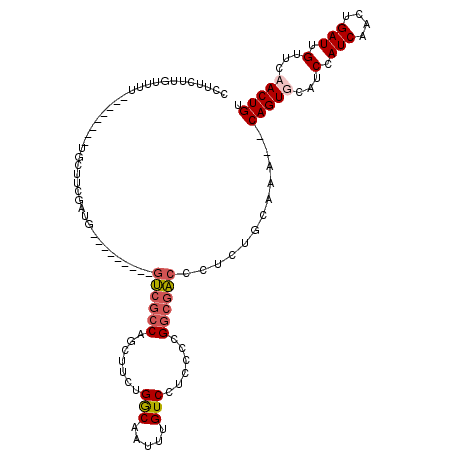
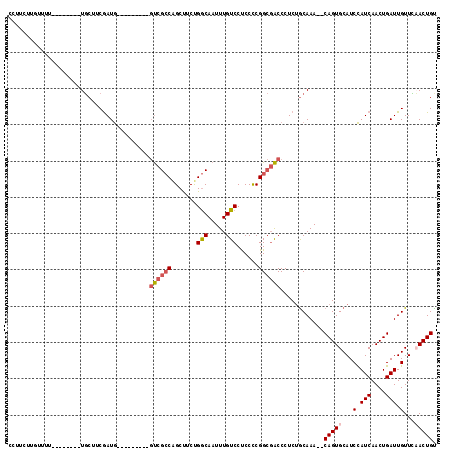
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:58:29 2006