
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,400,020 – 3,400,116 |
| Length | 96 |
| Max. P | 0.992294 |

| Location | 3,400,020 – 3,400,116 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 80.86 |
| Mean single sequence MFE | -23.00 |
| Consensus MFE | -15.70 |
| Energy contribution | -15.20 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.68 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992294 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
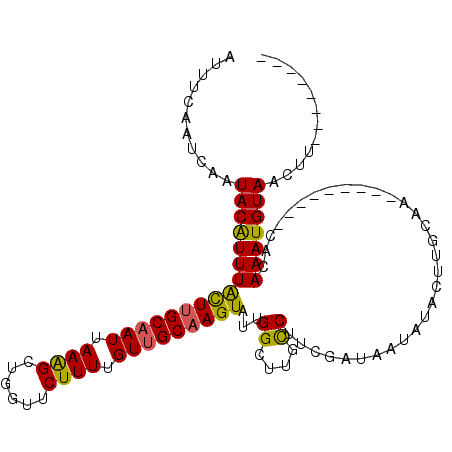
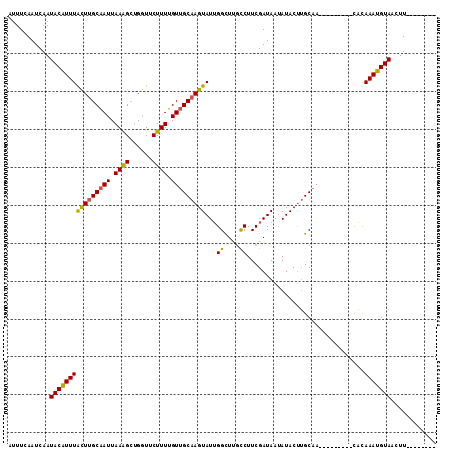
>2L_DroMel_CAF1 3400020 96 + 22407834 AUUUCAAUCAAUACAUUUGCUUGCAAUUAAAGGUGGUUCUUUUGUUGCAAGUAUUGGUUCGCCUUCGAUAAUAUACUUGCAA---------CACAAAUGUAAUUU-------- ...........((((((((.........(((((....)))))((((((((((((....(((....)))....))))))))))---------))))))))))....-------- ( -26.20) >DroSec_CAF1 22454 96 + 1 AUUUCAAUCAAUACAUUUACUUGCAAUUAAAGCUGGUUCUUUUGUUGCAAGUAUUGGCUAGUCUUCGAUAAUAUACUUGCCA---------CACAAAUGUAACUU-------- ...........((((((((((((((((.((((......)))).)))))))))..((((.(((.(........).))).))))---------...)))))))....-------- ( -23.50) >DroSim_CAF1 22672 96 + 1 AUUUCAAUCAAUACAUUUACUUGCAAUUAAAGCUGGUUCUUUUGUUGCAAGUAUUGGCUUGCCUUCGAUAAUAUACUUGCAA---------CACAAAUGUAACUU-------- ...........((((((((((.((.......)).))).....((((((((((((.((....)).........))))))))))---------)).)))))))....-------- ( -22.90) >DroEre_CAF1 22509 113 + 1 AUUUCAAUCAAUACGUUUGUUCGCCAUUAAGGCUGGUUCUUUGGUUGCAAGUAUGGGCUUUCCUUCGAUAAAAUACCGGCAUACUACUACUUCUAAAUGUAACUUUGAUAUAU ......(((((((((((((............((((((...((((..(.((((....)))).)..))))......)))))).............))))))))...))))).... ( -19.41) >consensus AUUUCAAUCAAUACAUUUACUUGCAAUUAAAGCUGGUUCUUUUGUUGCAAGUAUUGGCUUGCCUUCGAUAAUAUACUUGCAA_________CACAAAUGUAACUU________ ...........((((((((((((((((.((((......)))).)))))))))...((....))...............................)))))))............ (-15.70 = -15.20 + -0.50)

| Location | 3,400,020 – 3,400,116 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 80.86 |
| Mean single sequence MFE | -22.40 |
| Consensus MFE | -11.46 |
| Energy contribution | -12.14 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.686452 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
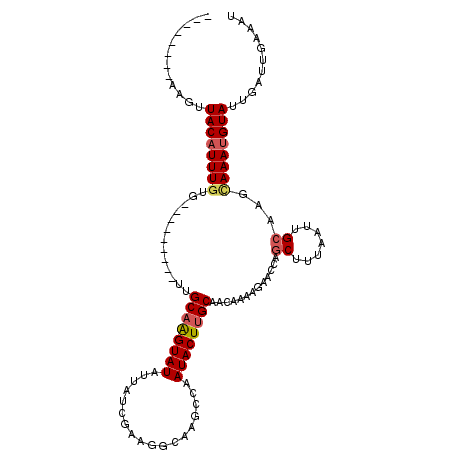
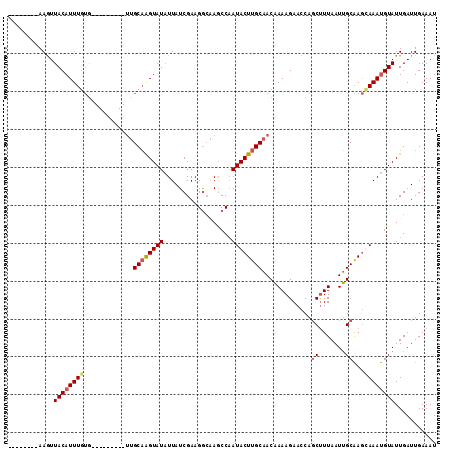
>2L_DroMel_CAF1 3400020 96 - 22407834 --------AAAUUACAUUUGUG---------UUGCAAGUAUAUUAUCGAAGGCGAACCAAUACUUGCAACAAAAGAACCACCUUUAAUUGCAAGCAAAUGUAUUGAUUGAAAU --------....((((((((((---------((((((((((....(((....)))....))))))))))).((((......))))........)))))))))........... ( -26.60) >DroSec_CAF1 22454 96 - 1 --------AAGUUACAUUUGUG---------UGGCAAGUAUAUUAUCGAAGACUAGCCAAUACUUGCAACAAAAGAACCAGCUUUAAUUGCAAGUAAAUGUAUUGAUUGAAAU --------....((((((....---------((((.(((...((....)).))).)))).(((((((((..((((......))))..)))))))))))))))........... ( -22.60) >DroSim_CAF1 22672 96 - 1 --------AAGUUACAUUUGUG---------UUGCAAGUAUAUUAUCGAAGGCAAGCCAAUACUUGCAACAAAAGAACCAGCUUUAAUUGCAAGUAAAUGUAUUGAUUGAAAU --------....((((((((((---------((((((((((.((....))((....)).)))))))))))).........((.......))...))))))))........... ( -22.20) >DroEre_CAF1 22509 113 - 1 AUAUAUCAAAGUUACAUUUAGAAGUAGUAGUAUGCCGGUAUUUUAUCGAAGGAAAGCCCAUACUUGCAACCAAAGAACCAGCCUUAAUGGCGAACAAACGUAUUGAUUGAAAU .....((((...(((........((.((((((((..(((.((((......)))).)))))))).))).))..........(((.....)))........)))....))))... ( -18.20) >consensus ________AAGUUACAUUUGUG_________UUGCAAGUAUAUUAUCGAAGGCAAGCCAAUACUUGCAACAAAAGAACCAGCUUUAAUUGCAAGCAAAUGUAUUGAUUGAAAU ............((((((((.............((((((((..................)))))))).............((.......))...))))))))........... (-11.46 = -12.14 + 0.69)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:58:24 2006