| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,248,305 – 3,248,415 |
| Length | 110 |
| Max. P | 0.968242 |

| Location | 3,248,305 – 3,248,415 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 84.69 |
| Mean single sequence MFE | -32.02 |
| Consensus MFE | -21.24 |
| Energy contribution | -21.05 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.98 |
| Structure conservation index | 0.66 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.968242 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3248305 110 - 22407834 AACUUUUCAACUUAUGCGUU-UGCUACUCUGGC--GCAGAAUAUUUU-GGCGAGUAUUCAUUCAGGUUUAUGCCAGCACAGUGGGUGGCUUAUGCAUAUGGCAUAUAUAUAUAC ..........(.(((((((.-.(((((((..(.--((.((((((((.-...)))))))).....(((....))).)).)...)))))))..))))))).).............. ( -32.30) >DroSim_CAF1 85515 106 - 1 AACUUUUCAACUUAUGCGUU-UGCUACUCUGGC--GCAGAAUAUUUU-GGCGAGUGU----UCAGGUUUAUGCCAGCACAGUGGGUGGCUUAUGCAUAUAGCAUAUAUAUAUAU ............(((((((.-.(((((((..(.--((.((((((((.-...))))))----)).(((....))).)).)...)))))))..)))))))................ ( -31.90) >DroEre_CAF1 79129 93 - 1 AACUUUUCAACUUAUGCGUU-UGCUACUCUUGCUGGCAGAAUAUUUU--GGGAGUGU----UCAGGUUUAUGCC---ACAGUGGGUGGCUUAUGCAUAUAGU----A------- .........((((((((((.-.(((((((..((((...((((((((.--..))))))----)).(((....)))---.)))))))))))..))))))).)))----.------- ( -32.80) >DroYak_CAF1 78390 101 - 1 AACUUUCCAACUUAUGCGUUUUGCUACUCUGGC--GCAGAAUAUUUUUGGGGAGUGU----UCAGGUUUAUGCCAGCACAGUGGGUGGCUUAUGCAUAUACUAUAUA------- ............(((((((...(((((((..(.--((.((((((((.....))))))----)).(((....))).)).)...)))))))..))))))).........------- ( -31.10) >consensus AACUUUUCAACUUAUGCGUU_UGCUACUCUGGC__GCAGAAUAUUUU_GGCGAGUGU____UCAGGUUUAUGCCAGCACAGUGGGUGGCUUAUGCAUAUAGCAUAUA_______ ............(((((((...(((((((...........((((((.....)))))).......(((....)))........)))))))..)))))))................ (-21.24 = -21.05 + -0.19)

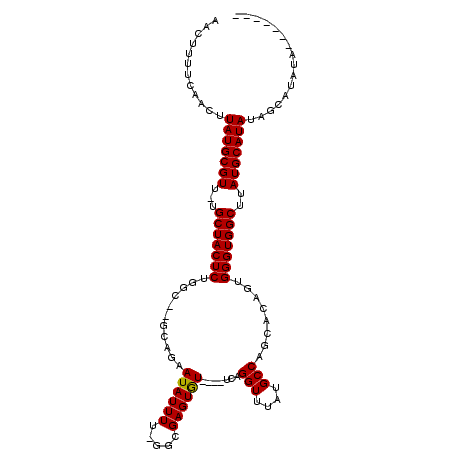
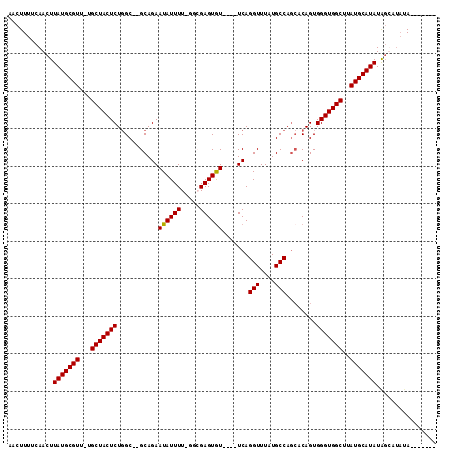
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:56:50 2006