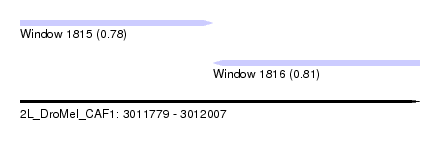

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,011,779 – 3,012,007 |
| Length | 228 |
| Max. P | 0.813389 |

| Location | 3,011,779 – 3,011,889 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 95.14 |
| Mean single sequence MFE | -24.51 |
| Consensus MFE | -23.66 |
| Energy contribution | -23.44 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.779448 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3011779 110 + 22407834 ACUUUUCCCUUUUUCGCUUUGCAUUUUCGUUUUUCCUUUUUGGCCAAUUUCAGGCCCCCCAAACGGGCUGAGAGCGAUUGCUAUUAUUUUGAGGUAAUUCGAUGAUUUAC .......((((.........(((...((((((((.......((((.......)))).(((....)))..)))))))).))).........))))................ ( -23.87) >DroSec_CAF1 19173 109 + 1 ACAUUUUCCUUUUUCGCUUUGCAUUUUCGUUUUUCCUU-UUGGCCAAUUUCAGGCCCCCCAAACGGGCUAAGAGCGAUUGCUGCUAUUUUGAGGUAAUUCGAUGAUUUAC .((((..((((....((...(((...((((((((....-..((((.......)))).(((....)))..)))))))).))).))......))))......))))...... ( -25.40) >DroEre_CAF1 19836 109 + 1 ACUUUUCCCUUCUUCGCUUUGCAUUUUCGUUUUUCCUU-UUGGCCAAUUUCAGGCCCCCCAAAAGGGCUGAGAGCGAUUGCUAUUAUUUUGAGGUAAUUCGAUGAUUUAC .......((((.........(((...((((((((((((-((((((.......))))....))))))...)))))))).))).........))))................ ( -24.27) >consensus ACUUUUCCCUUUUUCGCUUUGCAUUUUCGUUUUUCCUU_UUGGCCAAUUUCAGGCCCCCCAAACGGGCUGAGAGCGAUUGCUAUUAUUUUGAGGUAAUUCGAUGAUUUAC .......((((.........(((...((((((((.......((((.......)))).(((....)))..)))))))).))).........))))................ (-23.66 = -23.44 + -0.22)

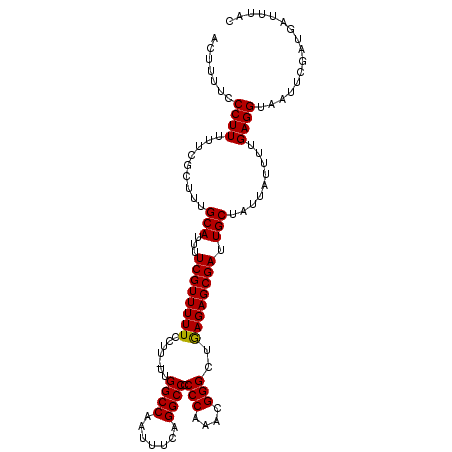
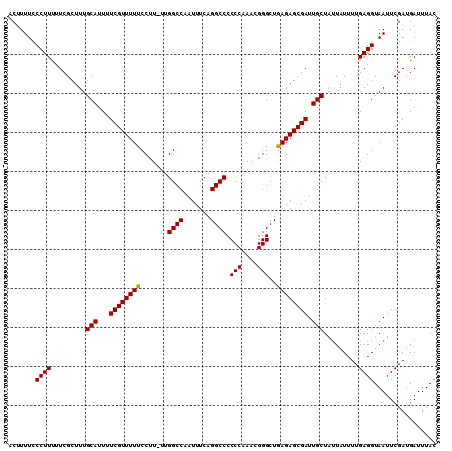
| Location | 3,011,889 – 3,012,007 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.97 |
| Mean single sequence MFE | -38.90 |
| Consensus MFE | -36.56 |
| Energy contribution | -36.12 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.813389 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 3011889 118 - 22407834 UUGUUGCACCCAGAGACGCAGACACCCGGGGUCUGAGGCACACACA--UAAAGCUCCGCUGGACAGCCAGUGGGAUCUGUGGCAUUCUGGAAAGAAAACUUACAAAACAAGGCAACGGUU (((((((..((((((.(.(((((.(....)))))).)((.((((.(--(.....((((((((....)))))))))).)))))).))))))........(((.......)))))))))).. ( -39.60) >DroSec_CAF1 19282 118 - 1 UUGUUGCACCCAGAGACGCAGACACCCGGGGUCUGAGGCACACACA--UAAAGCUCCGCUGGACAGCCAGUGGGAUCUGUGGCAUUCUGGAAAGAAAACUUACAAAACAAGGCAGCGGUU (((((((..((((((.(.(((((.(....)))))).)((.((((.(--(.....((((((((....)))))))))).)))))).))))))........(((.......)))))))))).. ( -39.60) >DroEre_CAF1 19945 120 - 1 UUGUUGCACCCAGAGACCCAGACACCCGGGGUCUGAAGCACACACAAAUACGGCUCCGCUGGACAGUCAGUGGGAUCUGUGGCAUUCUGGAAAGAAAACUUACAAAACAAGGCAACUGUU ..(((((..((((((((((.(.....).)))))....((.((((.......((..(((((((....)))))))..))))))))..)))))........(((.......)))))))).... ( -37.51) >consensus UUGUUGCACCCAGAGACGCAGACACCCGGGGUCUGAGGCACACACA__UAAAGCUCCGCUGGACAGCCAGUGGGAUCUGUGGCAUUCUGGAAAGAAAACUUACAAAACAAGGCAACGGUU ..(((((..((((((...(((((.(....))))))..((.((((..........((((((((....))))))))...)))))).))))))........(((.......)))))))).... (-36.56 = -36.12 + -0.44)

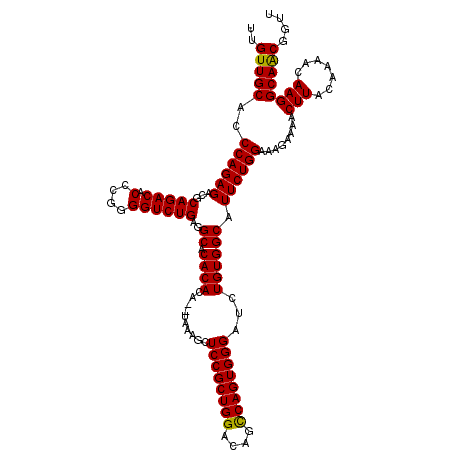
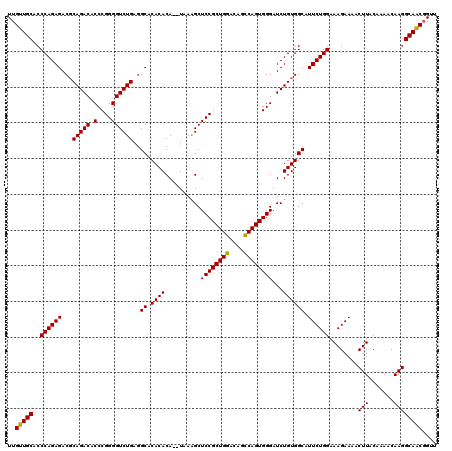
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:54:04 2006