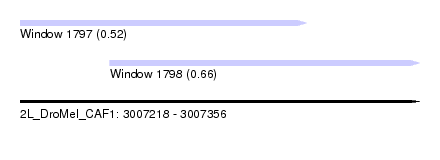

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 3,007,218 – 3,007,356 |
| Length | 138 |
| Max. P | 0.664498 |

| Location | 3,007,218 – 3,007,317 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.88 |
| Mean single sequence MFE | -32.50 |
| Consensus MFE | -22.67 |
| Energy contribution | -23.93 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.70 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.518438 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
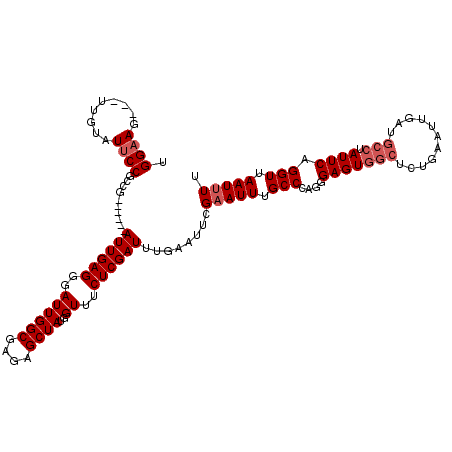
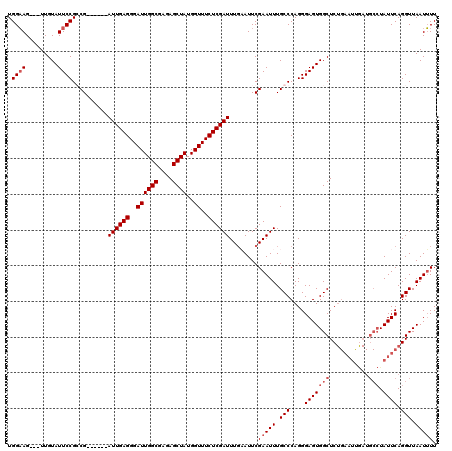
>2L_DroMel_CAF1 3007218 99 + 22407834 UGGAAG---UUGUAUUCCGGCG------AUUGAGGGAUUGGCGAGAGCUAUGGUUUCUCGAUUUGAAUUCGAAUUUGCCCAGGGAGUGGCUC------------UAUUCAGGUUAAUUUU ((((.(---(..(.((((((((------((((((..((((((....))))..))..)))))((((....)))).)))))..)))))..))))------------)).............. ( -28.40) >DroSec_CAF1 14723 111 + 1 UGGAAG---UUGUAUUCCGGCG------AUUGAGGGAUUGGCGAGAGCUAUGGUUUCUCGAUUUCAAUUCGAAUUUGCCCAGGGAGUGGCUCUGAAUUAAUGCCUAUUCAGGUUAAUUUU ..((((---(((((((((((((------((((((..((((((....))))..))..))))).(((.....))).)))))...))))))((.((((((........))))))))))))))) ( -31.90) >DroEre_CAF1 15242 111 + 1 UGGUAG---UUGUAUUCCGCCG------AUUGAGGGAUUGGCGAGAGCUAUGGUUUCUCGAUUUGUAUUCGAAUUUGCCCAGGGAGUGGCUCUGAAUCGAUGCCUAUUCAGGUUAAUAUU .((((.---(((.(((((((((------(((....))))))))(((((((((((...((((.......))))....)))......)))))))))))))))))))................ ( -32.50) >DroYak_CAF1 14407 120 + 1 UGGAAGUUGUUGUAUUCCGCCGAUUCCGAUUGAGGGAUUGGCGAGAGCUAUGGUUUCUCGAUUUGUAUUCGAAUUUGCCCAGGGAGUGGCUCUGAAUUGAUGCCUAUUCAGGUUAAUUUU .((((.........))))(((((((((......))))))))).(((((((((((...((((.......))))....)))......))))))))((((((..((((....)))))))))). ( -37.20) >consensus UGGAAG___UUGUAUUCCGCCG______AUUGAGGGAUUGGCGAGAGCUAUGGUUUCUCGAUUUGAAUUCGAAUUUGCCCAGGGAGUGGCUCUGAAUUGAUGCCUAUUCAGGUUAAUUUU .((((.........))))..........((((((..((((((....))))..))..))))))........(((((.(((....(((((((...........))).)))).))).))))). (-22.67 = -23.93 + 1.25)

| Location | 3,007,249 – 3,007,356 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.01 |
| Mean single sequence MFE | -34.82 |
| Consensus MFE | -24.44 |
| Energy contribution | -25.44 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.664498 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
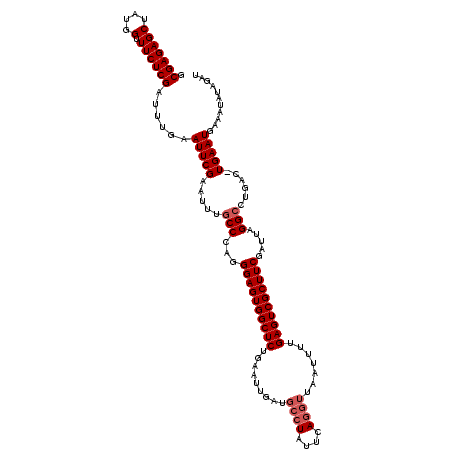
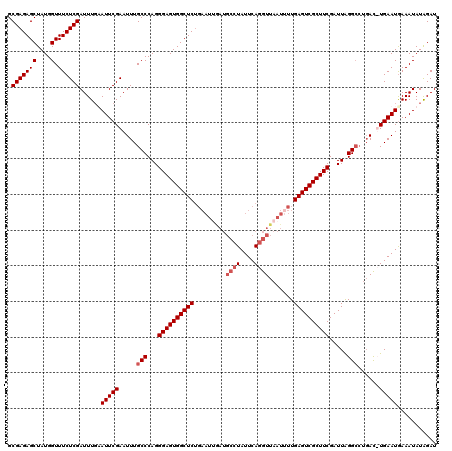
>2L_DroMel_CAF1 3007249 107 + 22407834 GCGAGAGCUAUGGUUUCUCGAUUUGAAUUCGAAUUUGCCCAGGGAGUGGCUC------------UAUUCAGGUUAAUUUUGAGUCGCUUCGAUUAGGCCCGAC-UGAAUGAAAUAUGGAU ((((((((....).)))))((((((....)))))).))(((.((((((((((------------................))))))))))..((((......)-)))........))).. ( -26.89) >DroSec_CAF1 14754 119 + 1 GCGAGAGCUAUGGUUUCUCGAUUUCAAUUCGAAUUUGCCCAGGGAGUGGCUCUGAAUUAAUGCCUAUUCAGGUUAAUUUUGAGUCGCUUCGAUUAGGCCUGAC-UGAAUGAAAUAUGGAU .(((((((....).))))))((((((.((((.....(((...((((((((((.((((((..((((....)))))))))).)))))))))).....))).....-))))))))))...... ( -36.80) >DroEre_CAF1 15273 120 + 1 GCGAGAGCUAUGGUUUCUCGAUUUGUAUUCGAAUUUGCCCAGGGAGUGGCUCUGAAUCGAUGCCUAUUCAGGUUAAUAUUGAGUCGCUUCGAUUAGGAGUGAUUUGAAUGAAAUAUAGAU .(((((((....).))))))(((((((((((.(((.(((....(((((((((......)).))).)))).))).))).(..(((((((((.....)))))))))..).)).))))))))) ( -34.50) >DroYak_CAF1 14447 120 + 1 GCGAGAGCUAUGGUUUCUCGAUUUGUAUUCGAAUUUGCCCAGGGAGUGGCUCUGAAUUGAUGCCUAUUCAGGUUAAUUUUGAGUCGCUUCGAUUGGGCCUCACUUGAAUAAAAUAUAGAU .(((((((....).))))))((((.(((((((....((((((((((((((((.((((((..((((....)))))))))).))))))))))..)))))).....)))))))))))...... ( -41.10) >consensus GCGAGAGCUAUGGUUUCUCGAUUUGAAUUCGAAUUUGCCCAGGGAGUGGCUCUGAAUUGAUGCCUAUUCAGGUUAAUUUUGAGUCGCUUCGAUUAGGCCUGAC_UGAAUGAAAUAUAGAU .(((((((....).))))))......(((((.....(((...((((((((((.........((((....)))).......)))))))))).....)))......)))))........... (-24.44 = -25.44 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:53:48 2006