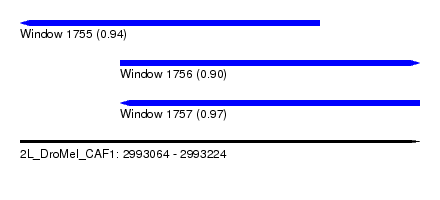

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,993,064 – 2,993,224 |
| Length | 160 |
| Max. P | 0.970882 |

| Location | 2,993,064 – 2,993,184 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.25 |
| Mean single sequence MFE | -43.17 |
| Consensus MFE | -41.08 |
| Energy contribution | -41.57 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.02 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938624 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
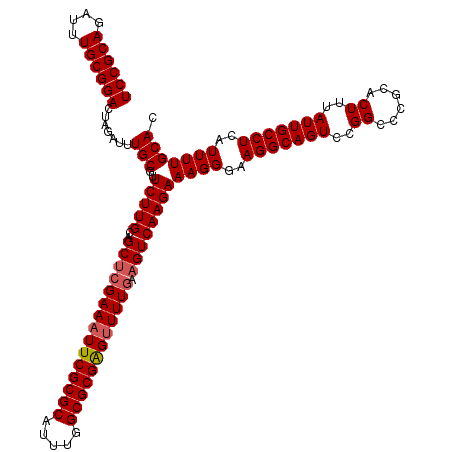
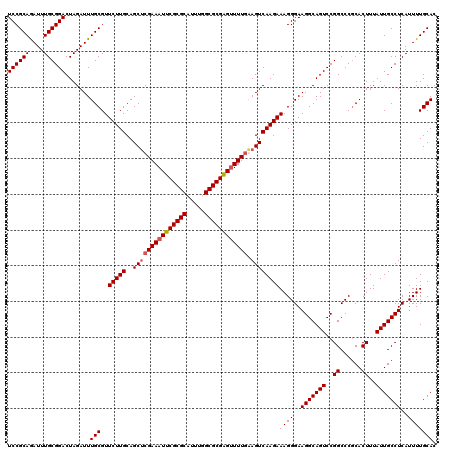
>2L_DroMel_CAF1 2993064 120 - 22407834 UCCGCAGAUUUGCGGACUAGAUUUGCGUUCUUGCAGCACGAAAUUCGCGCUUUUGGCGCGGGUUUUGAAGUCAAGAAAGGGAAGGCAGUCCGGCCCGCACUUUAUUGCCUCAUUUUGCAC ...(((((..((.((....((..(((.((((((.....((((((((((((.....))))))))))))....)))))).(((..((....))..))))))..))....)).)).))))).. ( -41.90) >DroSec_CAF1 4835 120 - 1 UCCGCAGAUUUGCGGACUAGAUUUGCGUUCUUGCAGCUCGAAAUUCGCGCAUUUGGCGCGAGUUUUGGAGUCAAGAAAGGGAAGGCAGUCCGGCCCGCACUUUAUUGCCUCAUUUUGCAC ...(((((..((.((....((..(((.((((((..(((((((((((((((.....))))))))))).)))))))))).(((..((....))..))))))..))....)).)).))))).. ( -45.30) >DroSim_CAF1 4809 120 - 1 UCCGCAGAUUUGCGGACUAGAUUUGCGUUCUUGCAGCUCGAAAUUCGCGCAUUUGGCGCGAGUUUUGGAGUCAAGAAAGGGAAGGCAGUCCGGCCCGCACUUUAUUGCCUCAUUUUGCAC ...(((((..((.((....((..(((.((((((..(((((((((((((((.....))))))))))).)))))))))).(((..((....))..))))))..))....)).)).))))).. ( -45.30) >DroYak_CAF1 4835 120 - 1 UCCGCAAAUUUGCGGACUAGAUUUGCGUUCUUGCAGCUCGAAAUUCGCGCUUUUGGCGCGGGAUUUUAAGUCAAGAAAGGGAAGGCAGUCCGGCCCACACUUUAUUGCCUCAUUUUGCAC ((((((....)))))).......(((..(((((..(((..((((((((((.....)))).))))))..))))))))((((..(((((((..((......))..)))))))..))))))). ( -40.20) >consensus UCCGCAGAUUUGCGGACUAGAUUUGCGUUCUUGCAGCUCGAAAUUCGCGCAUUUGGCGCGAGUUUUGAAGUCAAGAAAGGGAAGGCAGUCCGGCCCGCACUUUAUUGCCUCAUUUUGCAC ((((((....)))))).......(((..(((((..(((((((((((((((.....)))))))))))).))))))))((((..(((((((..((......))..)))))))..))))))). (-41.08 = -41.57 + 0.50)

| Location | 2,993,104 – 2,993,224 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -31.73 |
| Consensus MFE | -29.55 |
| Energy contribution | -29.55 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.901295 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 2993104 120 + 22407834 CCUUUCUUGACUUCAAAACCCGCGCCAAAAGCGCGAAUUUCGUGCUGCAAGAACGCAAAUCUAGUCCGCAAAUCUGCGGAAGUGUUGGUACUACCACUUAAUUGUAUUUGUAAUUGUAAU ...((((((.(..((.....(((((.....))))).......))..))))))).((((((.(((((((((....)))))(((((.((....)).))))).)))).))))))......... ( -28.90) >DroSec_CAF1 4875 120 + 1 CCUUUCUUGACUCCAAAACUCGCGCCAAAUGCGCGAAUUUCGAGCUGCAAGAACGCAAAUCUAGUCCGCAAAUCUGCGGAAGUGUUGGUACUACCACUUAAUUGUAUUUGUAAUUGUAAU ...((((((.(((..(((.((((((.....)))))).))).)))...)))))).((((((.(((((((((....)))))(((((.((....)).))))).)))).))))))......... ( -33.00) >DroSim_CAF1 4849 120 + 1 CCUUUCUUGACUCCAAAACUCGCGCCAAAUGCGCGAAUUUCGAGCUGCAAGAACGCAAAUCUAGUCCGCAAAUCUGCGGAAGUGUUGGUACUACCACUUAAUUGUAUUUGUAAUUGUAAU ...((((((.(((..(((.((((((.....)))))).))).)))...)))))).((((((.(((((((((....)))))(((((.((....)).))))).)))).))))))......... ( -33.00) >DroYak_CAF1 4875 120 + 1 CCUUUCUUGACUUAAAAUCCCGCGCCAAAAGCGCGAAUUUCGAGCUGCAAGAACGCAAAUCUAGUCCGCAAAUUUGCGGAAGUGUUGGUACUACCACUUAAUUGUAUUUGUAAUUGUAAU ...((((((.((..((((..(((((.....))))).))))..))...)))))).((((((.(((((((((....)))))(((((.((....)).))))).)))).))))))......... ( -32.00) >consensus CCUUUCUUGACUCCAAAACCCGCGCCAAAAGCGCGAAUUUCGAGCUGCAAGAACGCAAAUCUAGUCCGCAAAUCUGCGGAAGUGUUGGUACUACCACUUAAUUGUAUUUGUAAUUGUAAU ...((((((.(((.(((...(((((.....)))))..))).)))...)))))).((((((.(((((((((....)))))(((((.((....)).))))).)))).))))))......... (-29.55 = -29.55 + 0.00)

| Location | 2,993,104 – 2,993,224 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -36.03 |
| Consensus MFE | -34.71 |
| Energy contribution | -35.02 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.970882 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
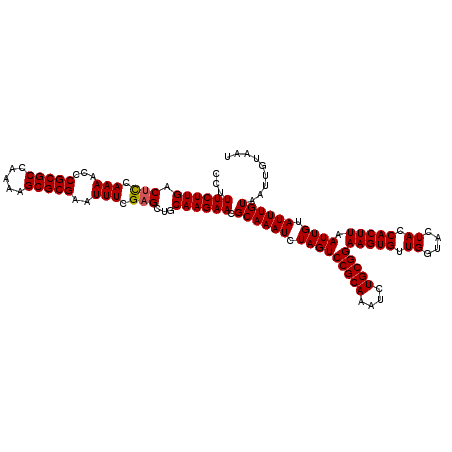
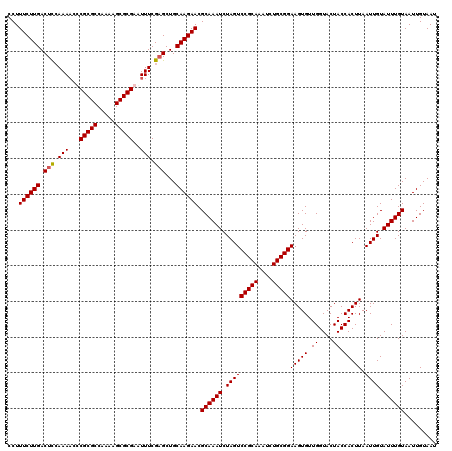
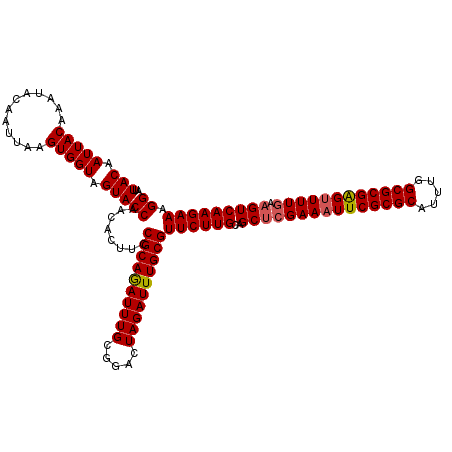
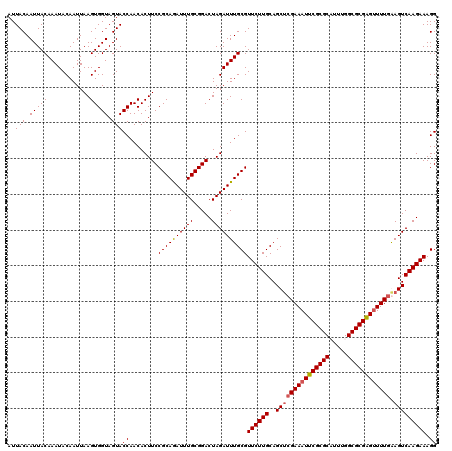
>2L_DroMel_CAF1 2993104 120 - 22407834 AUUACAAUUACAAAUACAAUUAAGUGGUAGUACCAACACUUCCGCAGAUUUGCGGACUAGAUUUGCGUUCUUGCAGCACGAAAUUCGCGCUUUUGGCGCGGGUUUUGAAGUCAAGAAAGG ..(((.(((((............))))).)))((......(((((......)))))((.((((((((....)))....((((((((((((.....))))))))))))))))).))...)) ( -34.90) >DroSec_CAF1 4875 120 - 1 AUUACAAUUACAAAUACAAUUAAGUGGUAGUACCAACACUUCCGCAGAUUUGCGGACUAGAUUUGCGUUCUUGCAGCUCGAAAUUCGCGCAUUUGGCGCGAGUUUUGGAGUCAAGAAAGG ..(((.(((((............))))).)))((........((((((((((.....))))))))))((((((..(((((((((((((((.....))))))))))).)))))))))).)) ( -38.00) >DroSim_CAF1 4849 120 - 1 AUUACAAUUACAAAUACAAUUAAGUGGUAGUACCAACACUUCCGCAGAUUUGCGGACUAGAUUUGCGUUCUUGCAGCUCGAAAUUCGCGCAUUUGGCGCGAGUUUUGGAGUCAAGAAAGG ..(((.(((((............))))).)))((........((((((((((.....))))))))))((((((..(((((((((((((((.....))))))))))).)))))))))).)) ( -38.00) >DroYak_CAF1 4875 120 - 1 AUUACAAUUACAAAUACAAUUAAGUGGUAGUACCAACACUUCCGCAAAUUUGCGGACUAGAUUUGCGUUCUUGCAGCUCGAAAUUCGCGCUUUUGGCGCGGGAUUUUAAGUCAAGAAAGG ..(((.(((((............))))).)))((........((((((((((.....))))))))))((((((..(((..((((((((((.....)))).))))))..))))))))).)) ( -33.20) >consensus AUUACAAUUACAAAUACAAUUAAGUGGUAGUACCAACACUUCCGCAGAUUUGCGGACUAGAUUUGCGUUCUUGCAGCUCGAAAUUCGCGCAUUUGGCGCGAGUUUUGAAGUCAAGAAAGG ..(((.(((((............))))).)))((........((((((((((.....))))))))))((((((..(((((((((((((((.....)))))))))))).))))))))).)) (-34.71 = -35.02 + 0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:53:08 2006