| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,924,105 – 2,924,204 |
| Length | 99 |
| Max. P | 0.998707 |

| Location | 2,924,105 – 2,924,204 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.09 |
| Mean single sequence MFE | -26.79 |
| Consensus MFE | -15.74 |
| Energy contribution | -15.80 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.59 |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.986981 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
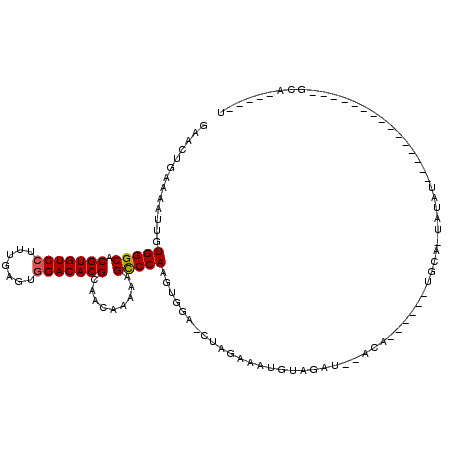
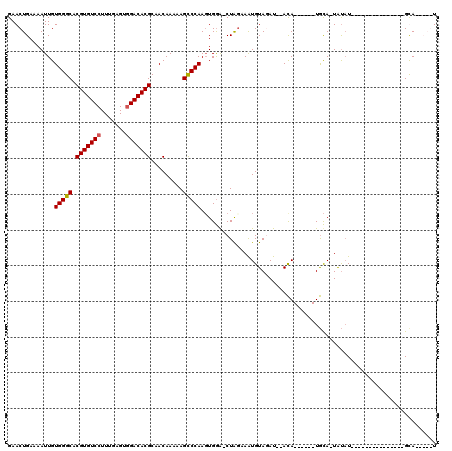
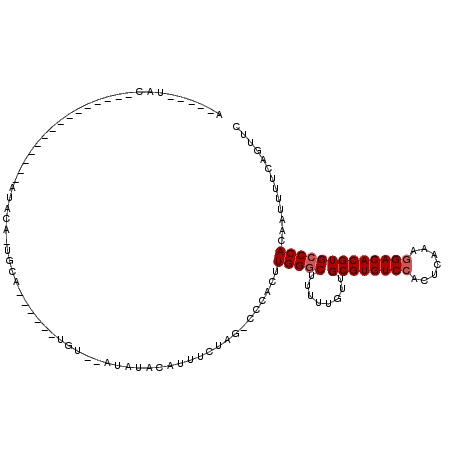
>2L_DroMel_CAF1 2924105 99 + 22407834 GAACUGAAAAUUGUGGGCACGUGUCCUUUGAGUGGACACGCAACAAAAAGCCCAACUGAAUCUAGAAGUGUAUAUAUACA------UGUAGUAUAU---------------GCAUAUAGU ..((((.......(((((.(((((((.......))))))).........))))).............((((((((((...------....))))))---------------)))).)))) ( -27.20) >DroSec_CAF1 42294 90 + 1 GAACUGAAAAUUGUGGGCACGUGUCCUUUGAGUGGACACGCAACAAAAAGUCCAAGUGGG-CUAGAAAUGUAGAU--ACA------UGCA-UGUAU---------------GUA-----U ...(((....((((..((..((((((.......)))))))).))))..(((((....)))-)).......)))((--(((------((..-..)))---------------)))-----) ( -24.80) >DroSim_CAF1 43141 90 + 1 GAACUGAAAAUUGUGGGCACGUGUCCUUUGAGUGGACACGCAACAAAAAGCCCAAGUGGA-CUAGAAAUGUAGAU--ACA------UGCA-UGUAU---------------GUA-----U ...(((.......(((((.(((((((.......))))))).........)))))......-.)))........((--(((------((..-..)))---------------)))-----) ( -23.44) >DroEre_CAF1 43251 116 + 1 GAACUGAAAAUUGUGGGCACGUGUCCUUUGAGGCGACACGCAACAAAAAGCCCAAGUGGG-CUGCCAAUGUAUAU--AUAUGUACAUACA-UACUUAUAUUUUAAAAAGUCGCAUAUAGU ..((((.....(((((((.((((((((....)).))))))........(((((....)))-)))))...(((((.--.(((((....)))-))..)))))..........))))..)))) ( -31.70) >consensus GAACUGAAAAUUGUGGGCACGUGUCCUUUGAGUGGACACGCAACAAAAAGCCCAAGUGGA_CUAGAAAUGUAGAU__ACA______UGCA_UAUAU_______________GCA_____U .............(((((.(((((((.......))))))).........))))).................................................................. (-15.74 = -15.80 + 0.06)


| Location | 2,924,105 – 2,924,204 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.09 |
| Mean single sequence MFE | -24.27 |
| Consensus MFE | -16.42 |
| Energy contribution | -16.92 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.68 |
| SVM decision value | 3.19 |
| SVM RNA-class probability | 0.998707 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
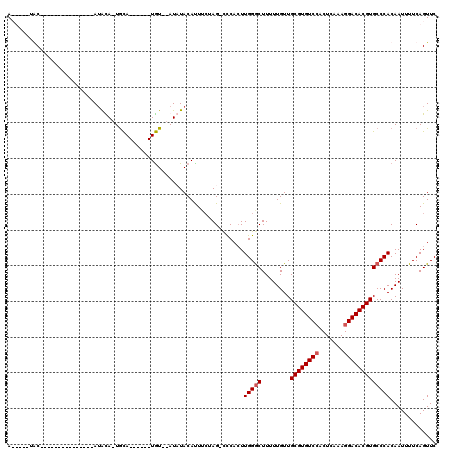
>2L_DroMel_CAF1 2924105 99 - 22407834 ACUAUAUGC---------------AUAUACUACA------UGUAUAUAUACACUUCUAGAUUCAGUUGGGCUUUUUGUUGCGUGUCCACUCAAAGGACACGUGCCCACAAUUUUCAGUUC ..(((((((---------------((.......)------))))))))..........((......(((((........((((((((.......)))))))))))))......))..... ( -26.60) >DroSec_CAF1 42294 90 - 1 A-----UAC---------------AUACA-UGCA------UGU--AUCUACAUUUCUAG-CCCACUUGGACUUUUUGUUGCGUGUCCACUCAAAGGACACGUGCCCACAAUUUUCAGUUC (-----(((---------------((...-...)------)))--))............-.......(((((..(((((((((((((.......)))))))))...)))).....))))) ( -18.40) >DroSim_CAF1 43141 90 - 1 A-----UAC---------------AUACA-UGCA------UGU--AUCUACAUUUCUAG-UCCACUUGGGCUUUUUGUUGCGUGUCCACUCAAAGGACACGUGCCCACAAUUUUCAGUUC (-----(((---------------((...-...)------)))--))............-......(((((........((((((((.......)))))))))))))............. ( -22.10) >DroEre_CAF1 43251 116 - 1 ACUAUAUGCGACUUUUUAAAAUAUAAGUA-UGUAUGUACAUAU--AUAUACAUUGGCAG-CCCACUUGGGCUUUUUGUUGCGUGUCGCCUCAAAGGACACGUGCCCACAAUUUUCAGUUC .........((((.............(((-((((((....)))--))))))...(((((-(((....))))).......(((((((.........))))))))))..........)))). ( -30.00) >consensus A_____UAC_______________AUACA_UGCA______UGU__AUAUACAUUUCUAG_CCCACUUGGGCUUUUUGUUGCGUGUCCACUCAAAGGACACGUGCCCACAAUUUUCAGUUC ..................................................................(((((........((((((((.......)))))))))))))............. (-16.42 = -16.92 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:51:46 2006